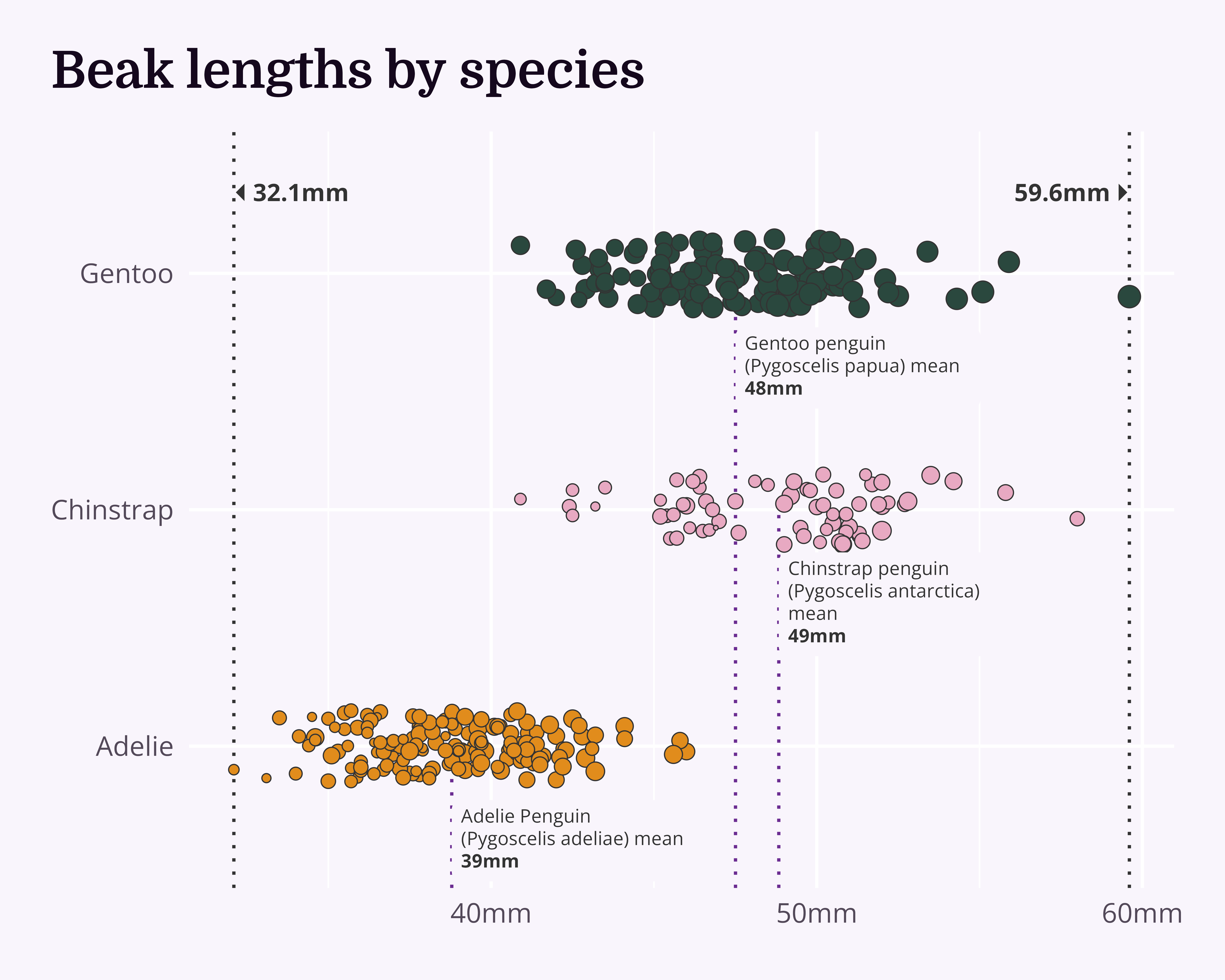
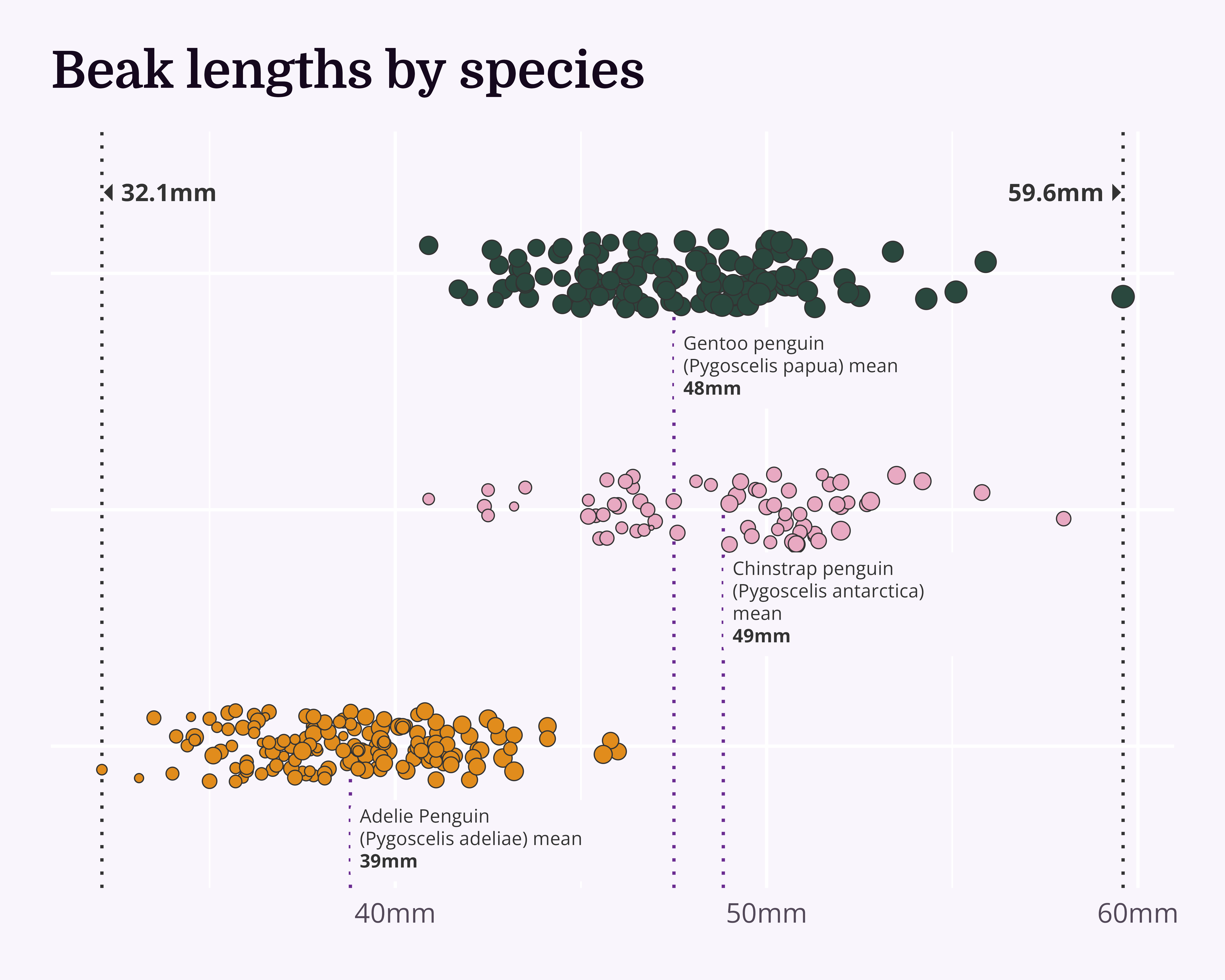
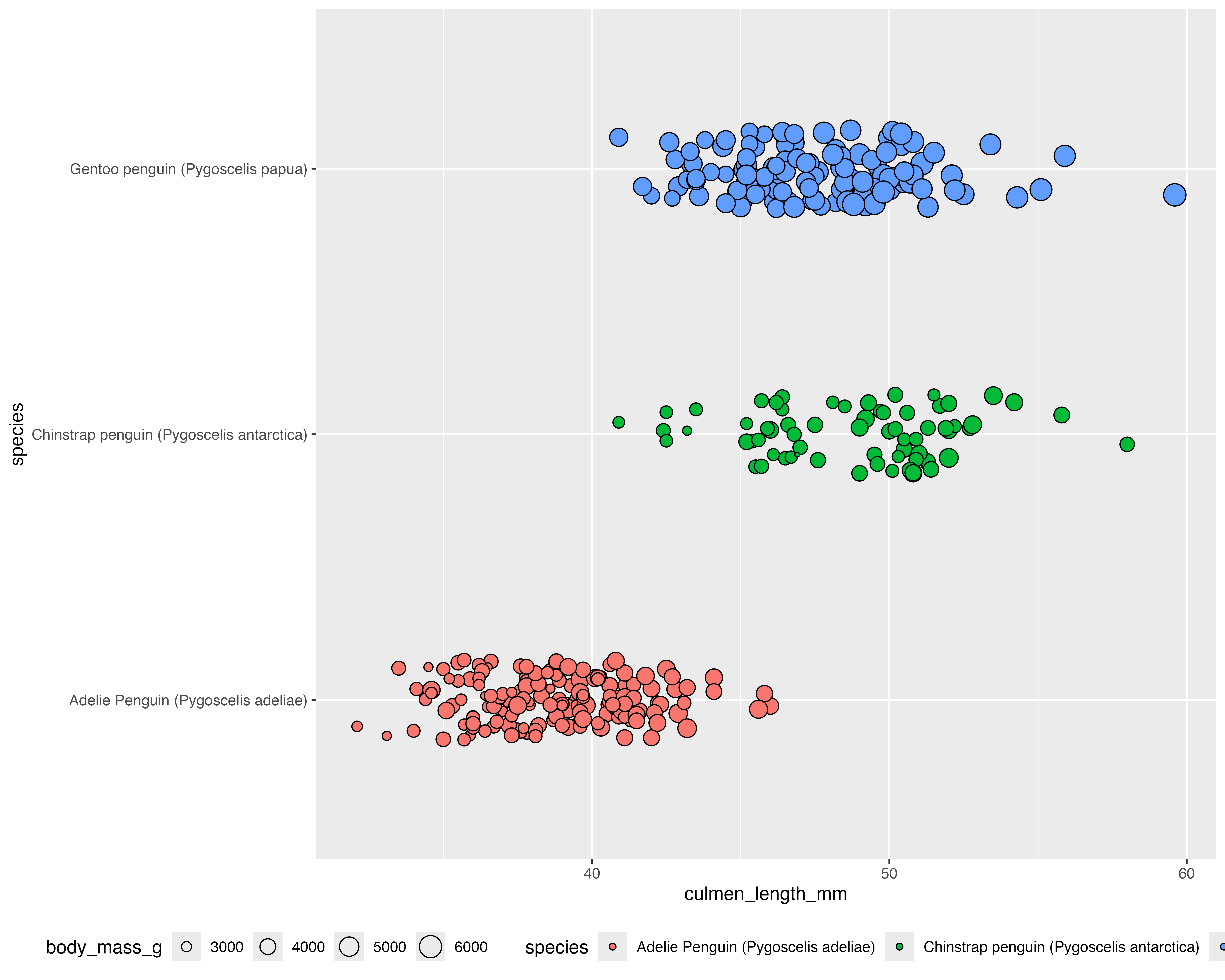
Make it your own
Tips & trick for stand-out dataviz with R and ggplot2
Cara R Thompson, PhD
26th March 2026
Hello 👋
👩 Cara Thompson
👩💻 Love for patterns in music & language, and a fascination with the human brain %>%
Psychology PhD %>%
Analysis of postgraduate medical examinations %>%
Data Visualisation Consultant
💙 Helping others maximise the impact of their expertise
Find out more: cararthompson.com/about
How did I end up here?
How did I end up here?
How did I end up here?
Why dataviz?
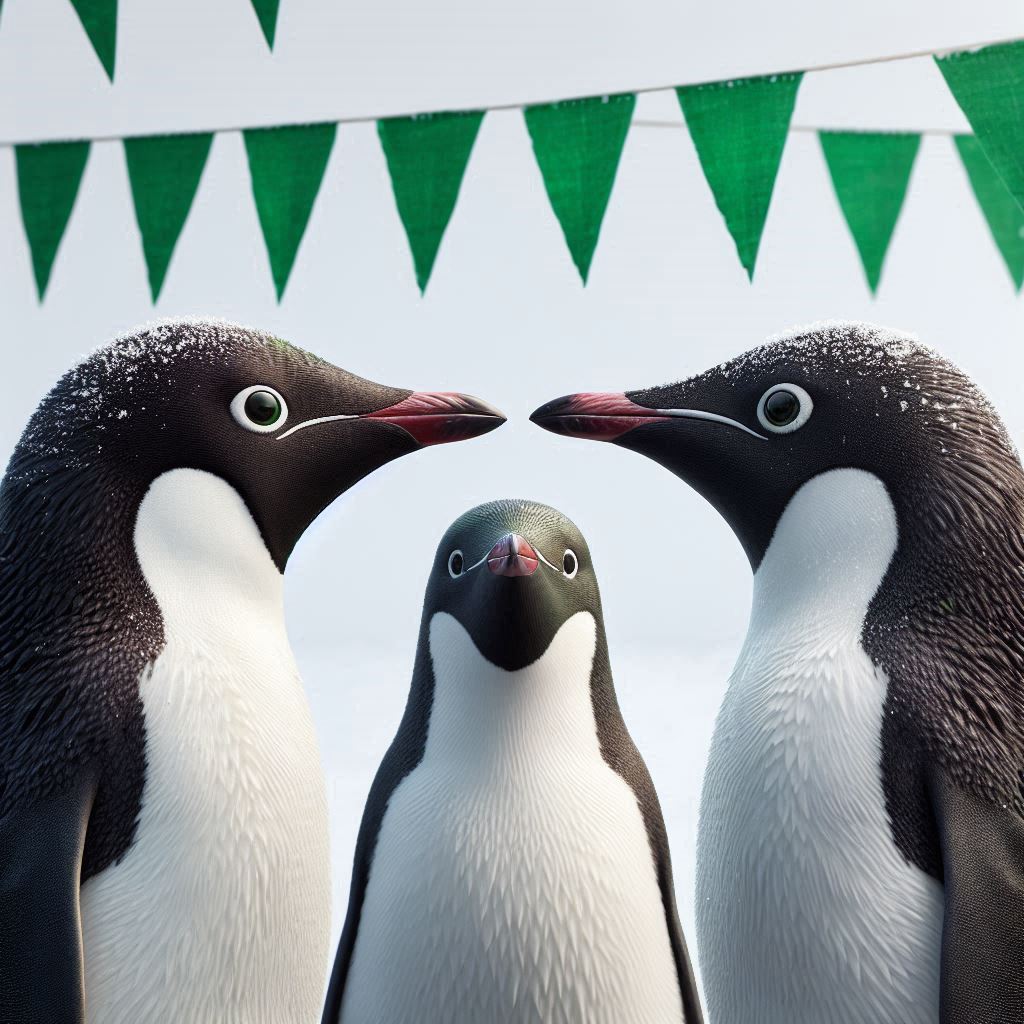
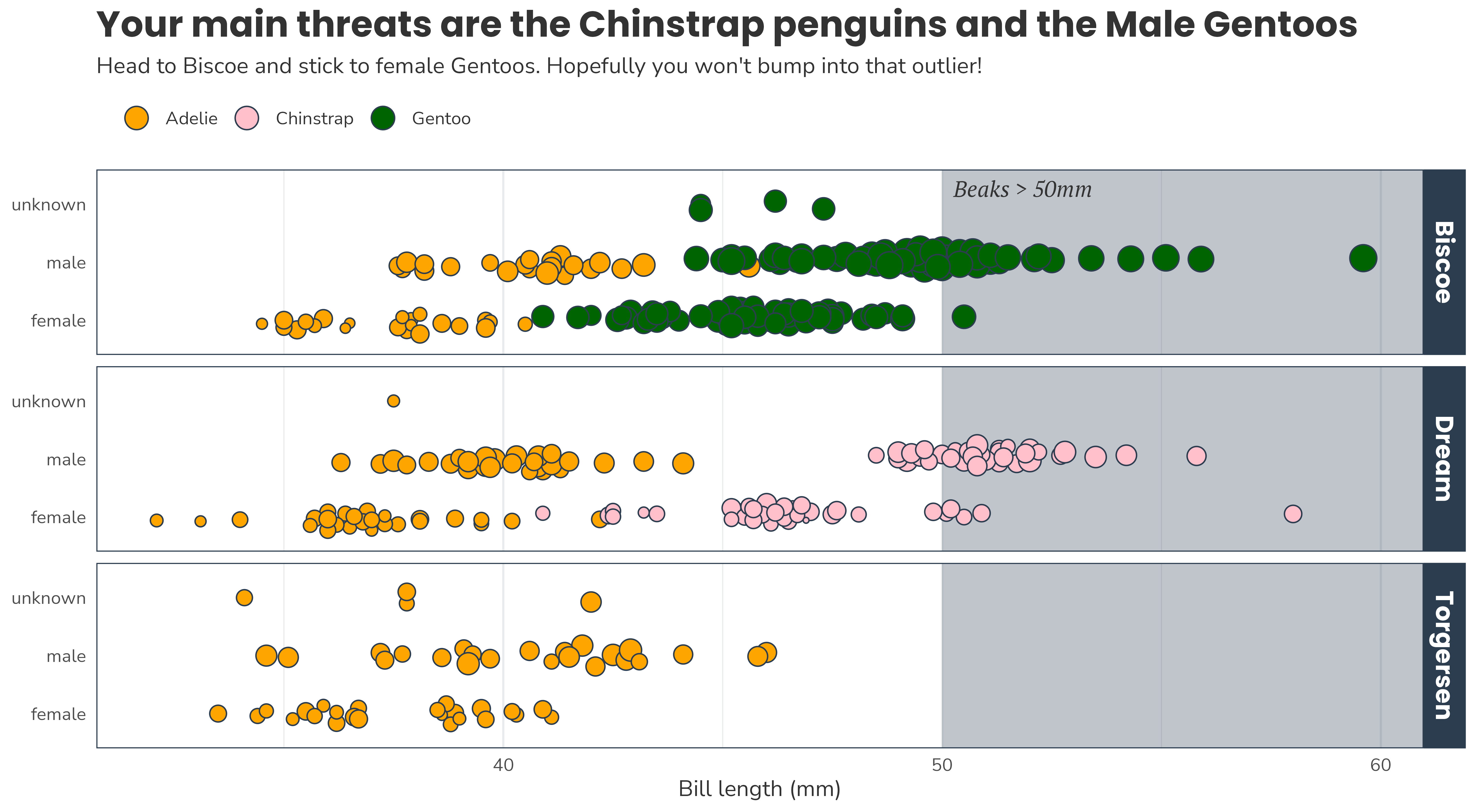
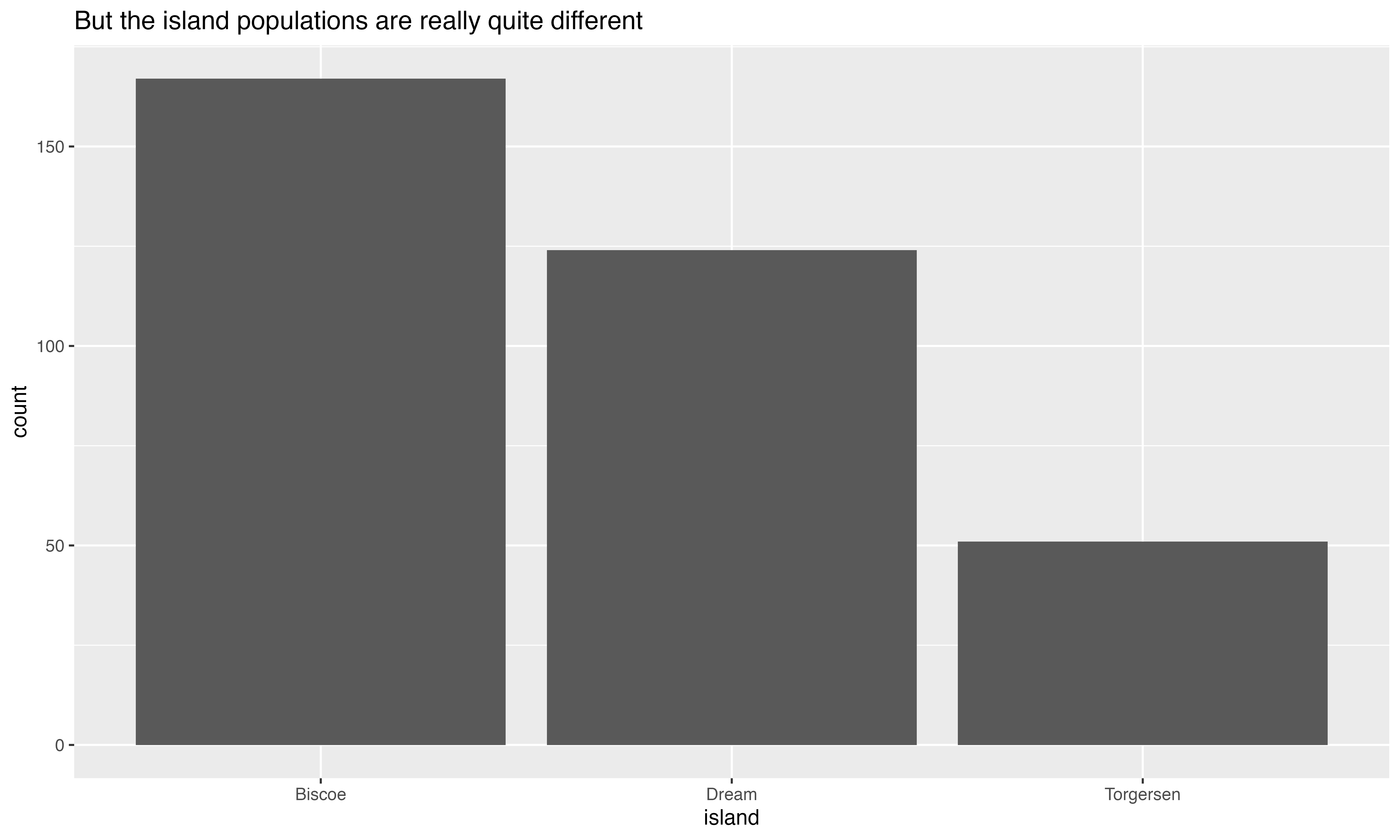
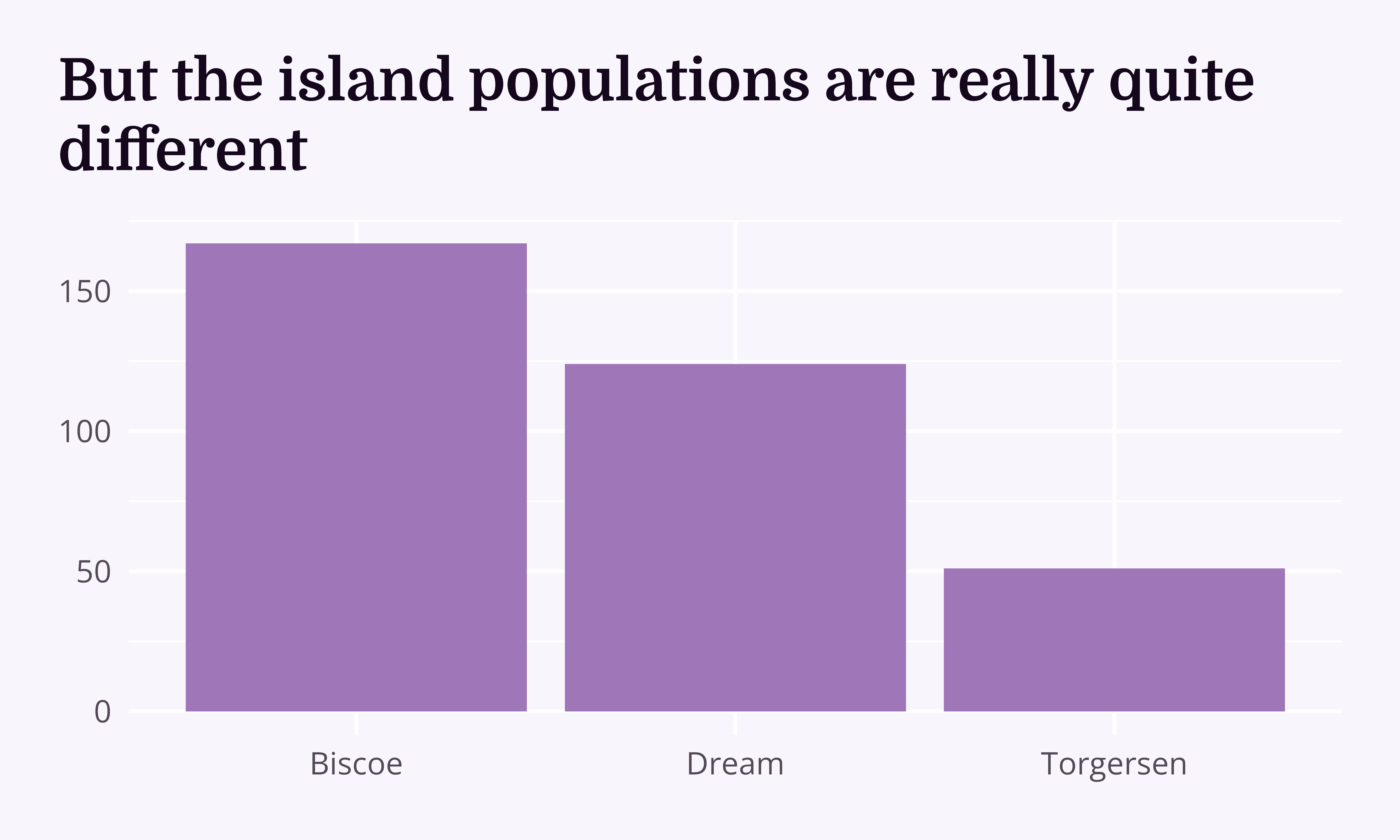
I’m planning an expedition to the islands where the Palmer Penguins live. I need to see at least 2 penguin species on my trip, but I can only go to one island.
I’m terrified of penguins with long beaks.
Help me plan my trip. You can only show me one slide.
Why dataviz?
“We have collected data about 344 penguins who live on Biscoe, Dream and Torgersen. On Biscoe, we have 44 Adelie penguins (F = 22, M = 22) and 124 Gentoos (F = 58, M = 61, Unknown = 5). On Dream, we have 56 Adelie penguins (F = 27, M = 28, Unknown = 1) and 68 Chinstrap penguins (M = F = 34). On Torgersen, we only have Adelies (F = 24, M = 23, Unknown = 5).
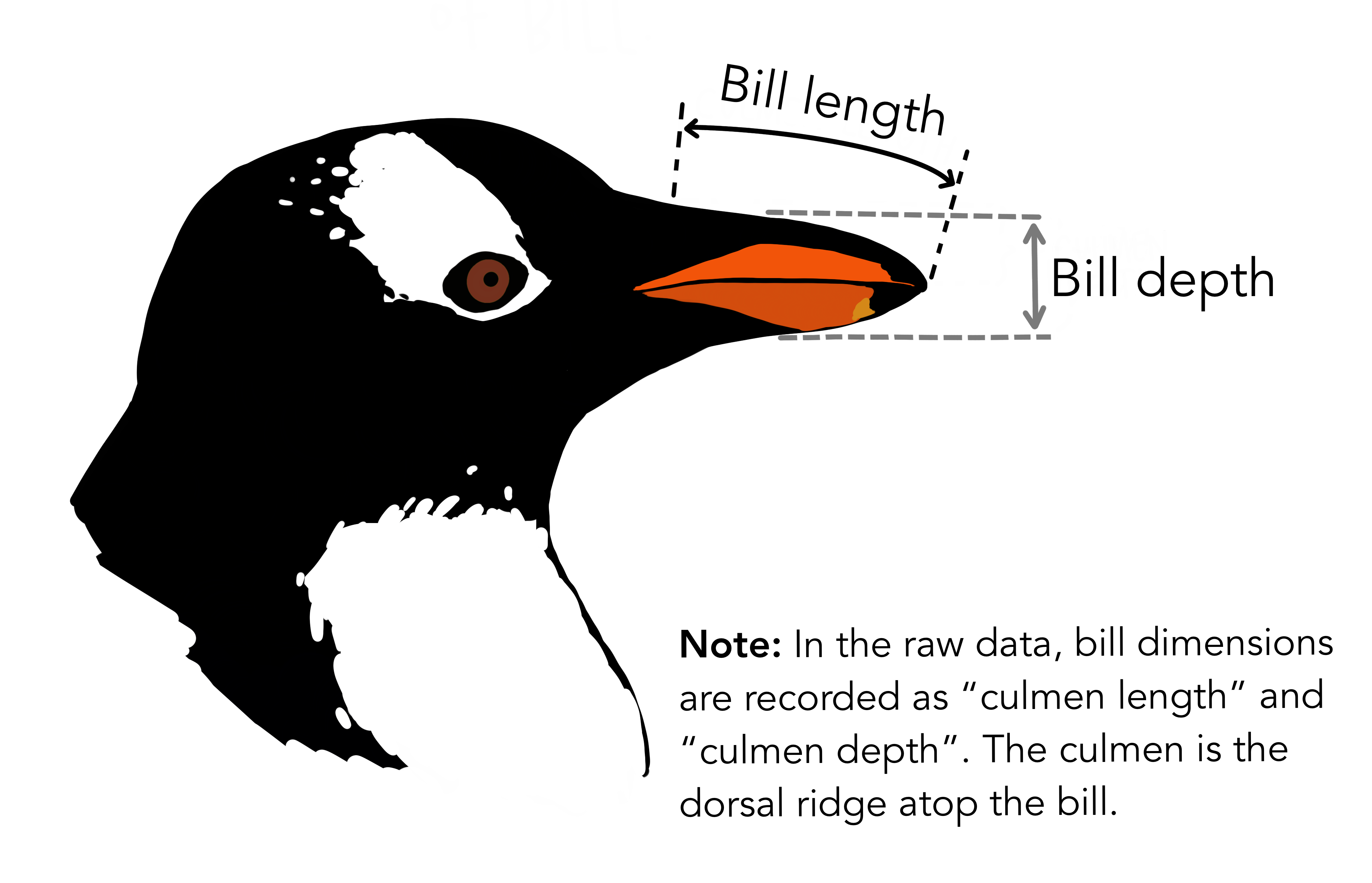
The average beak lengths of the species are as follows:
- Adelie: F: 37.55 mm, M: 40.59mm
- Chinstrap: F: 46.57mm, M: 51.09mm
- Gentoo: F: 45.59mm, M: 49.47mm
Have a nice trip!”
Why dataviz?
“Does this help?”
Why dataviz?
“Ok, let me reorganise it a bit”
Why dataviz?
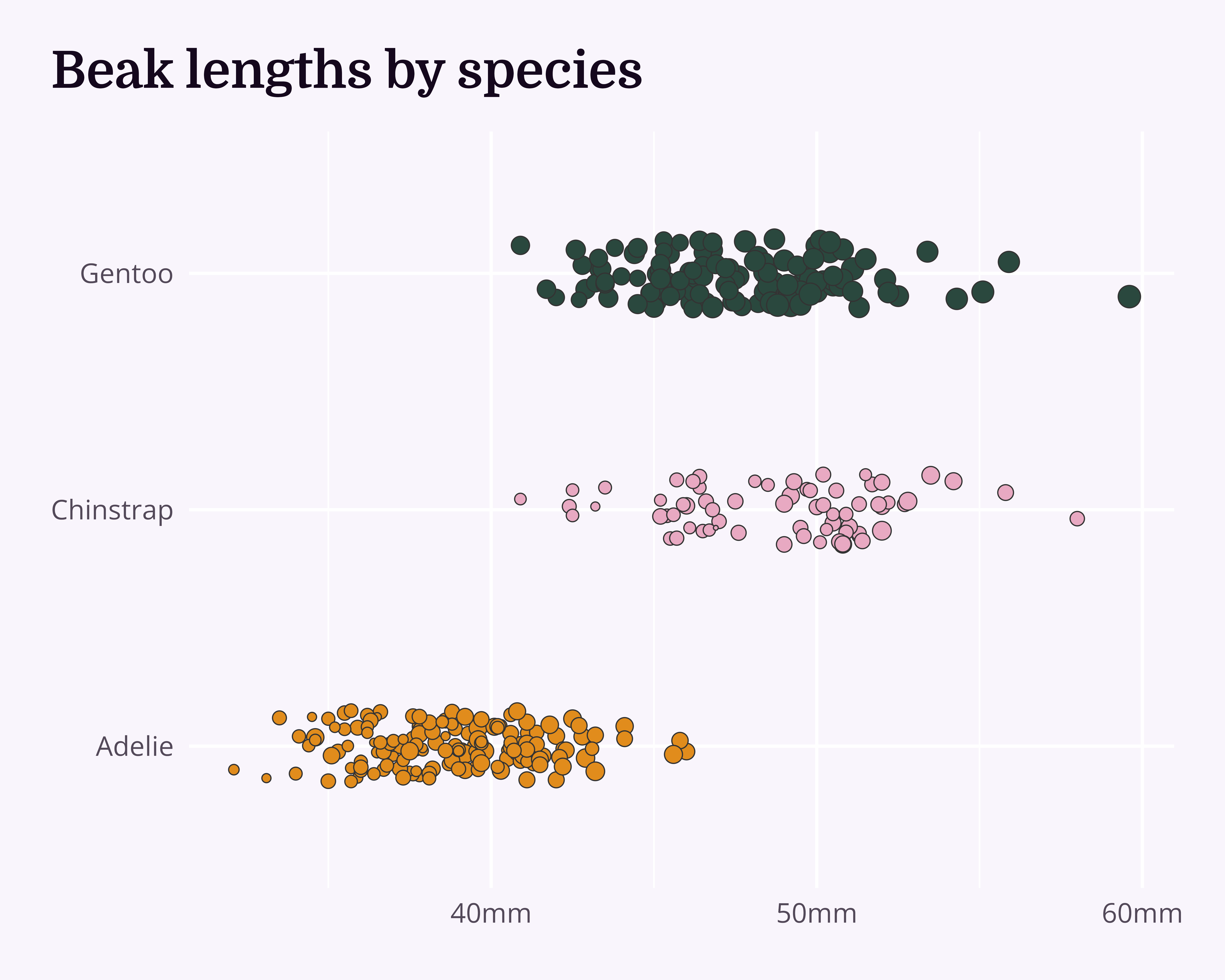
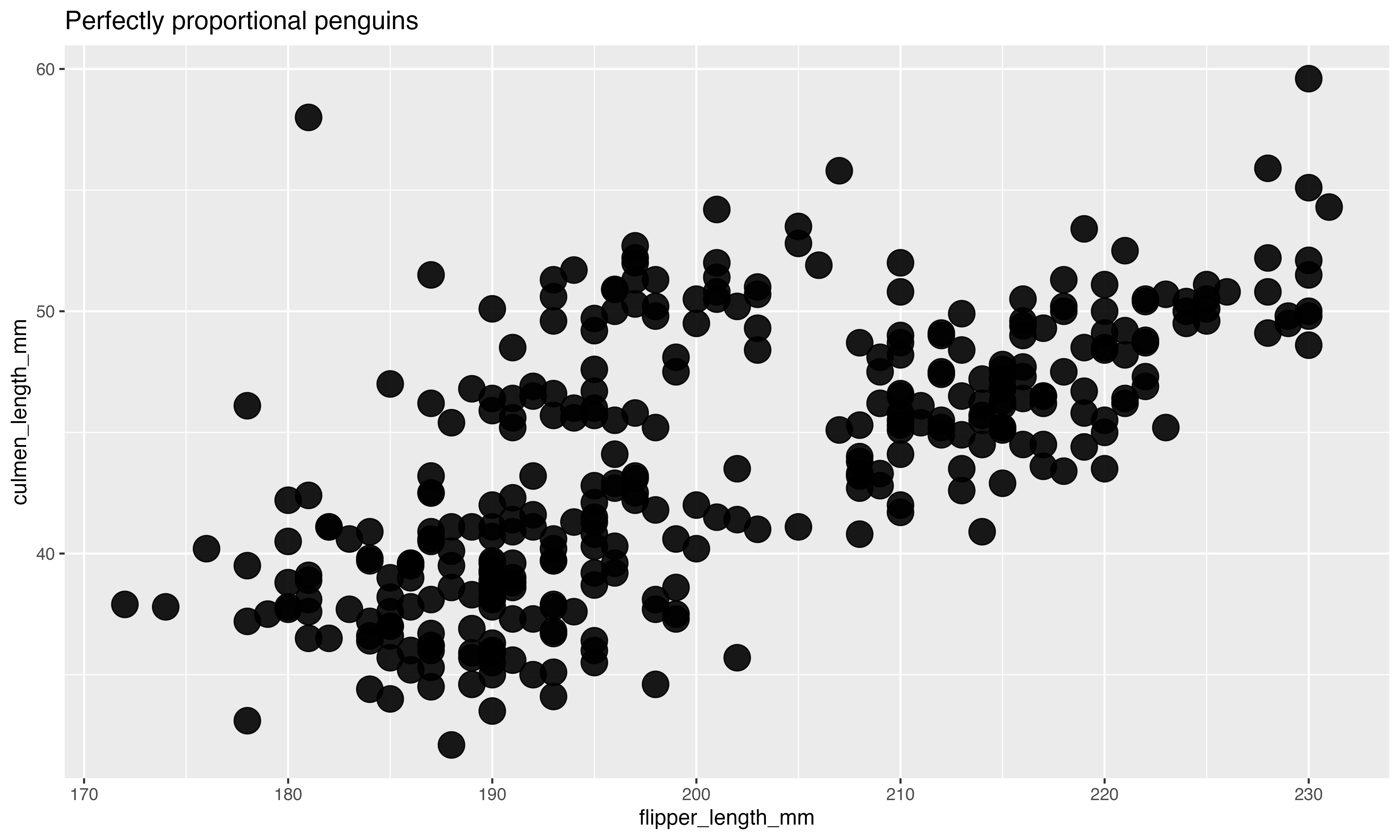
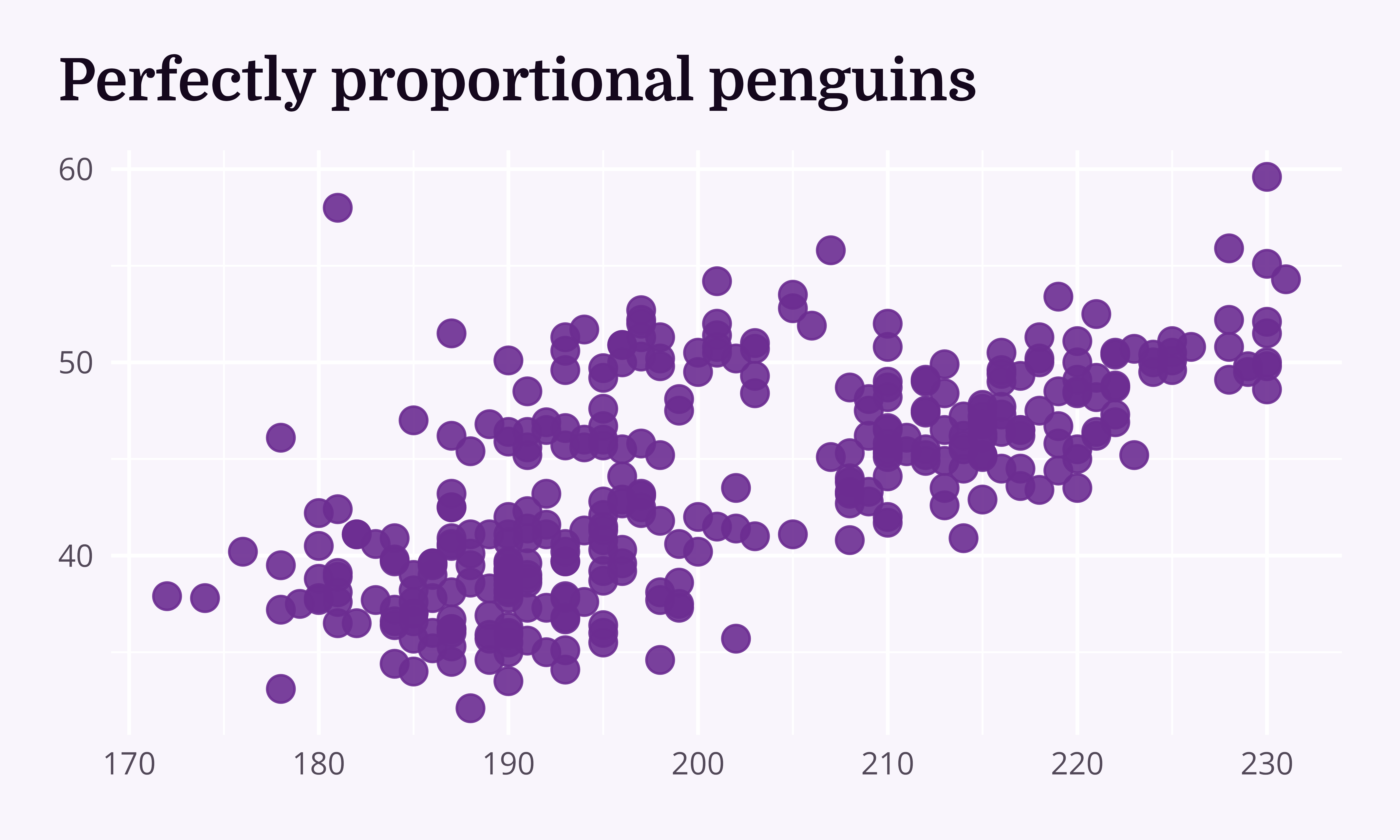
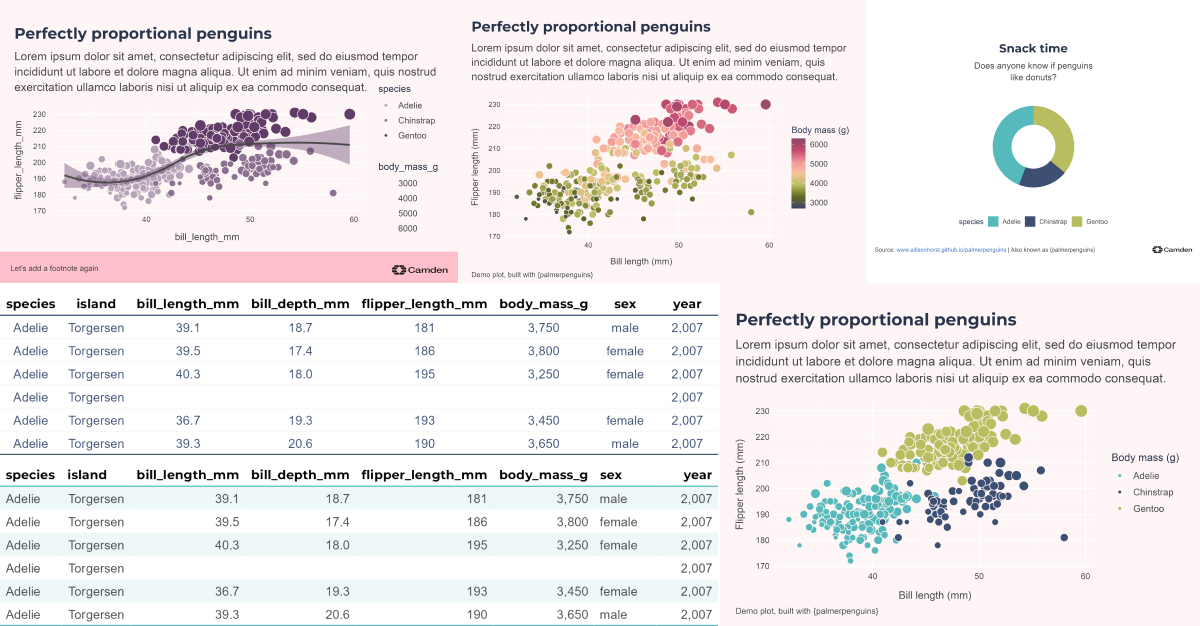
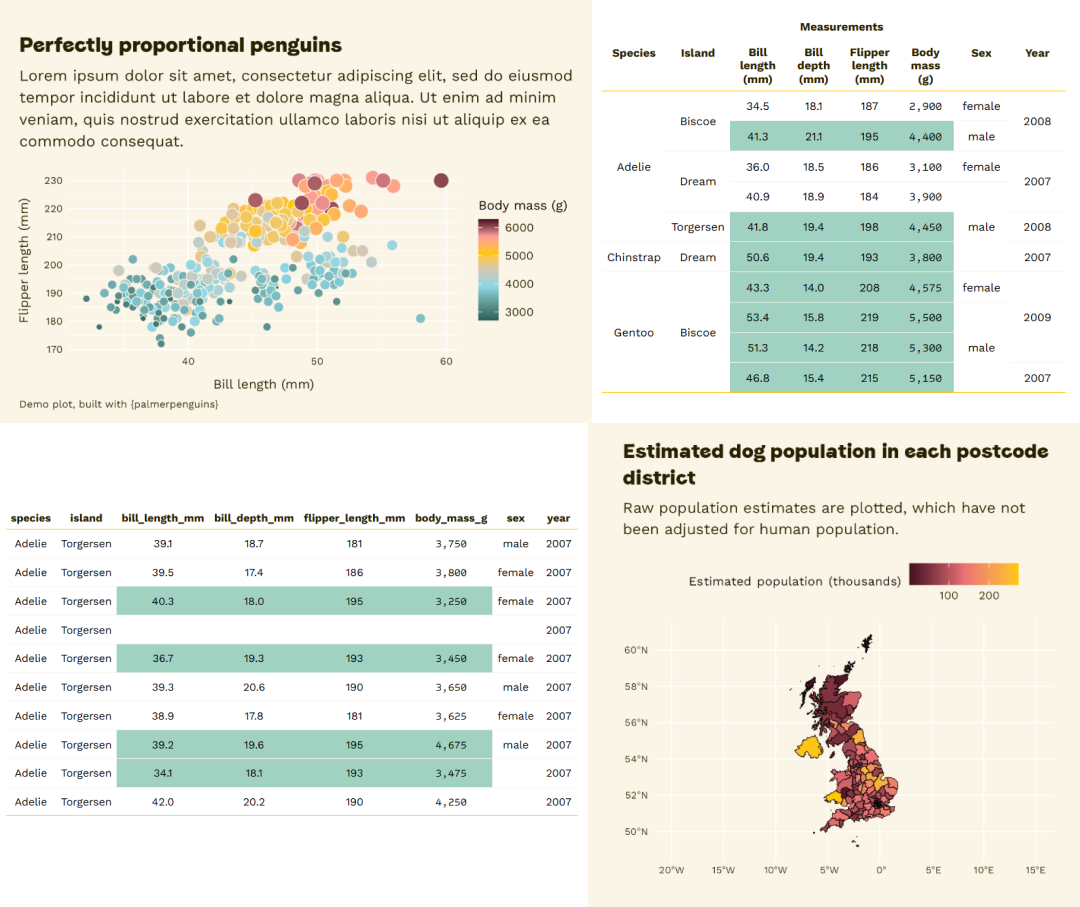
“Look, the data really speaks for itself… Here you go!”
Why dataviz better?
Visualise
What’s the best way to visualise the story?
Chester Zoo is welcoming some new penguins from Edinburgh Zoo. The keepers are a bit nervous about how the penguins will all get on.
They get quite competitive within each species about their beak lengths.
If we can see which penguin is which, even better!
Illustration by Allison Horst
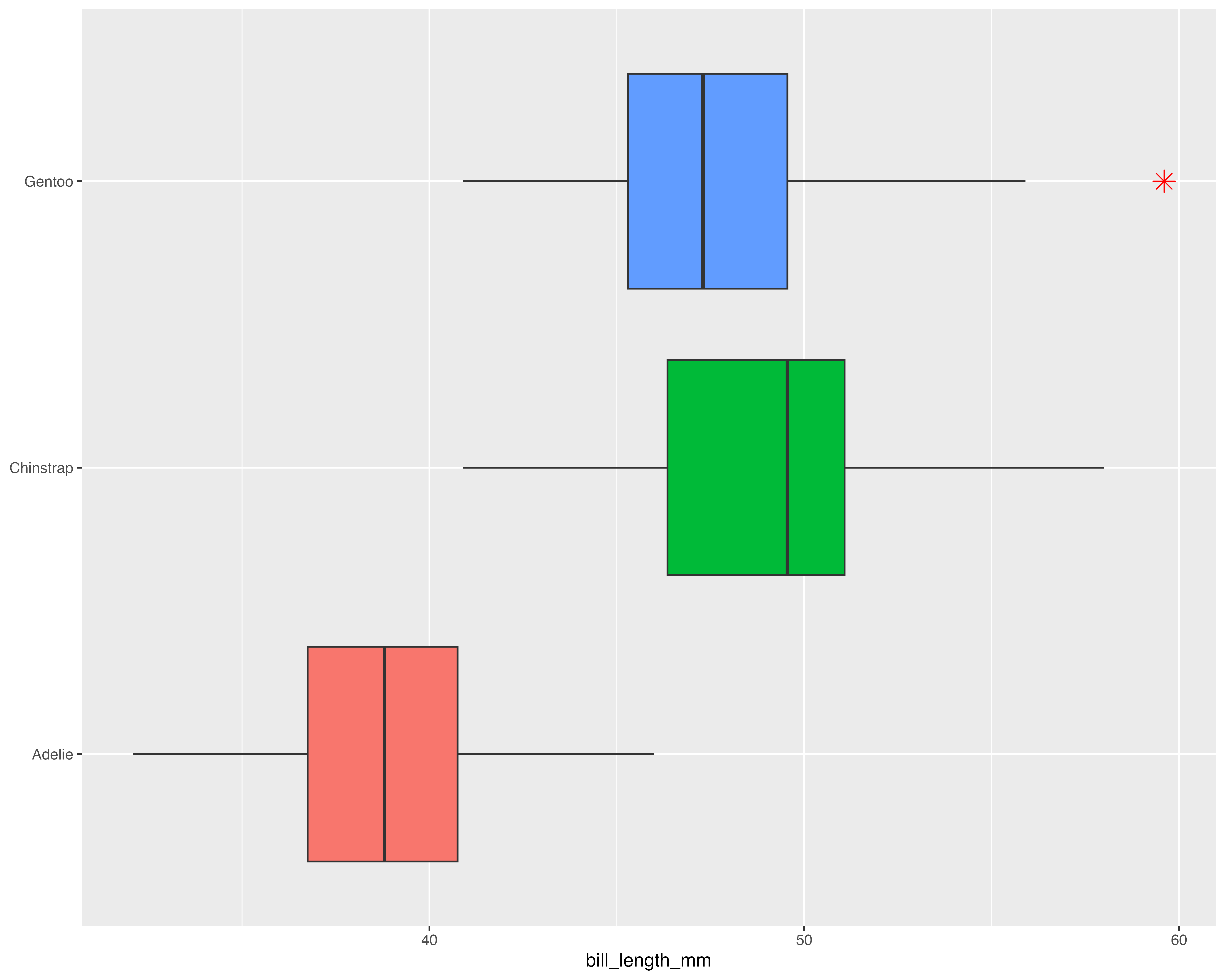
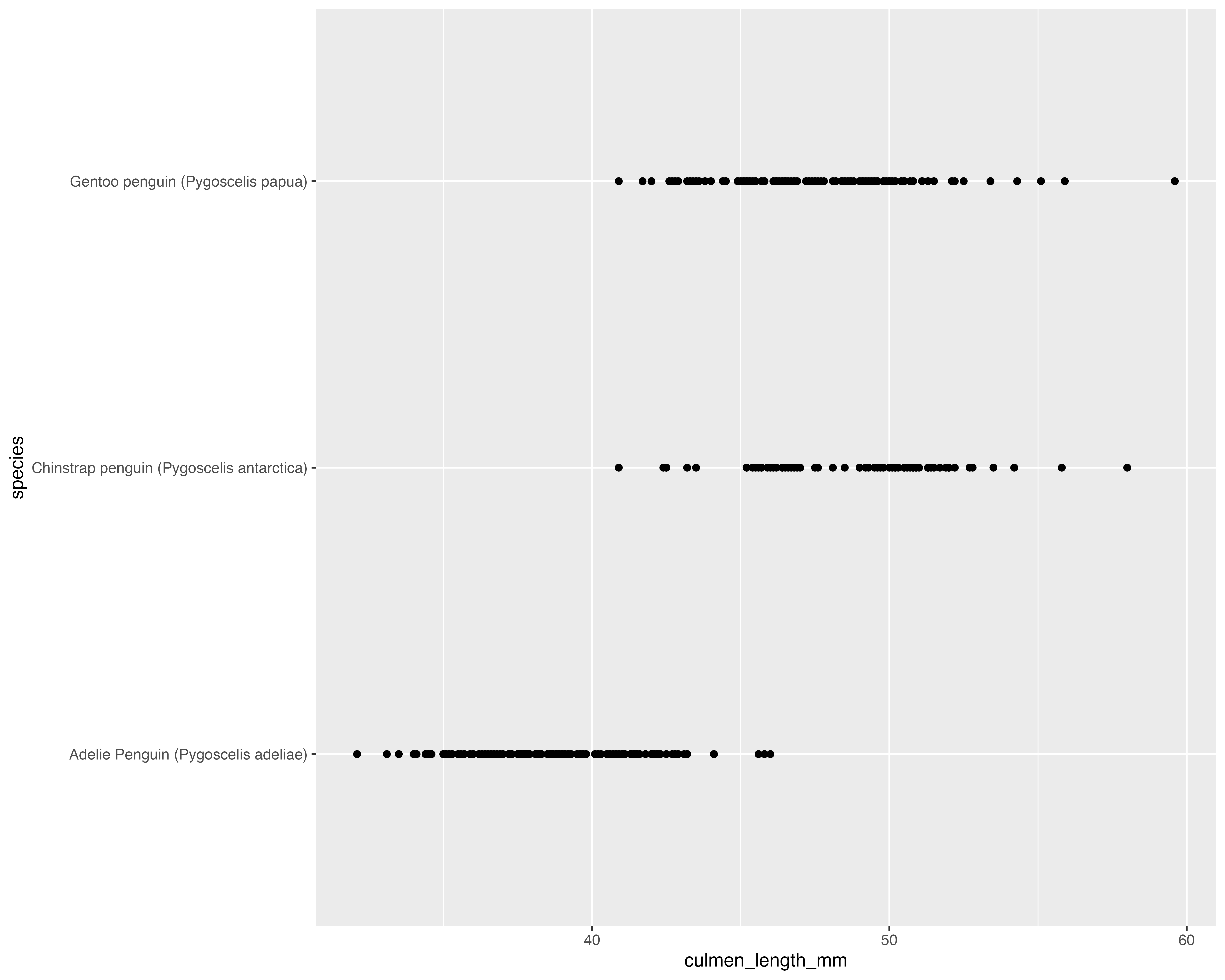
What’s the best way to visualise the story?
What’s the best way to visualise the story?
What’s the best way to visualise the story?
What’s the best way to visualise the story?
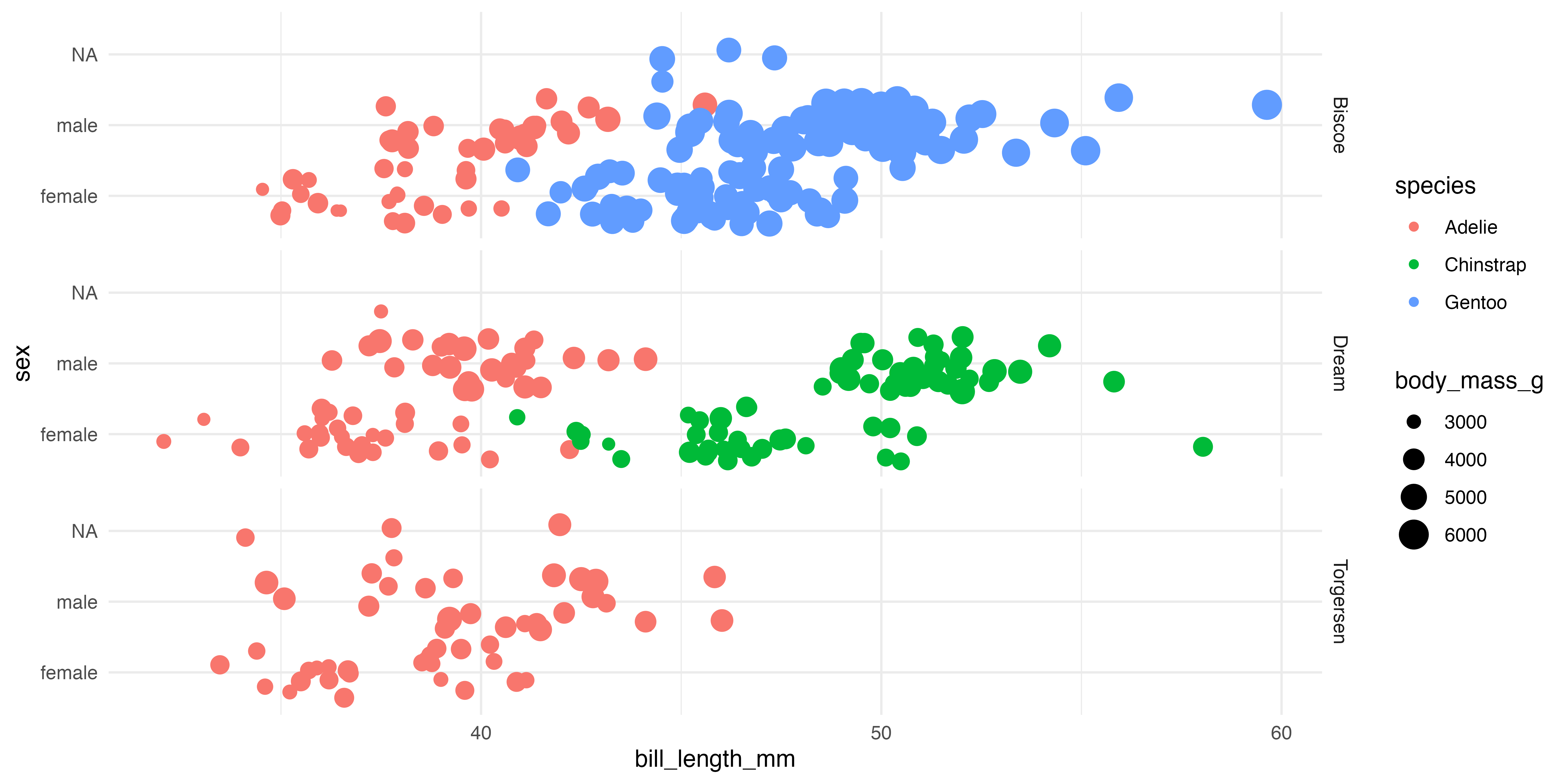
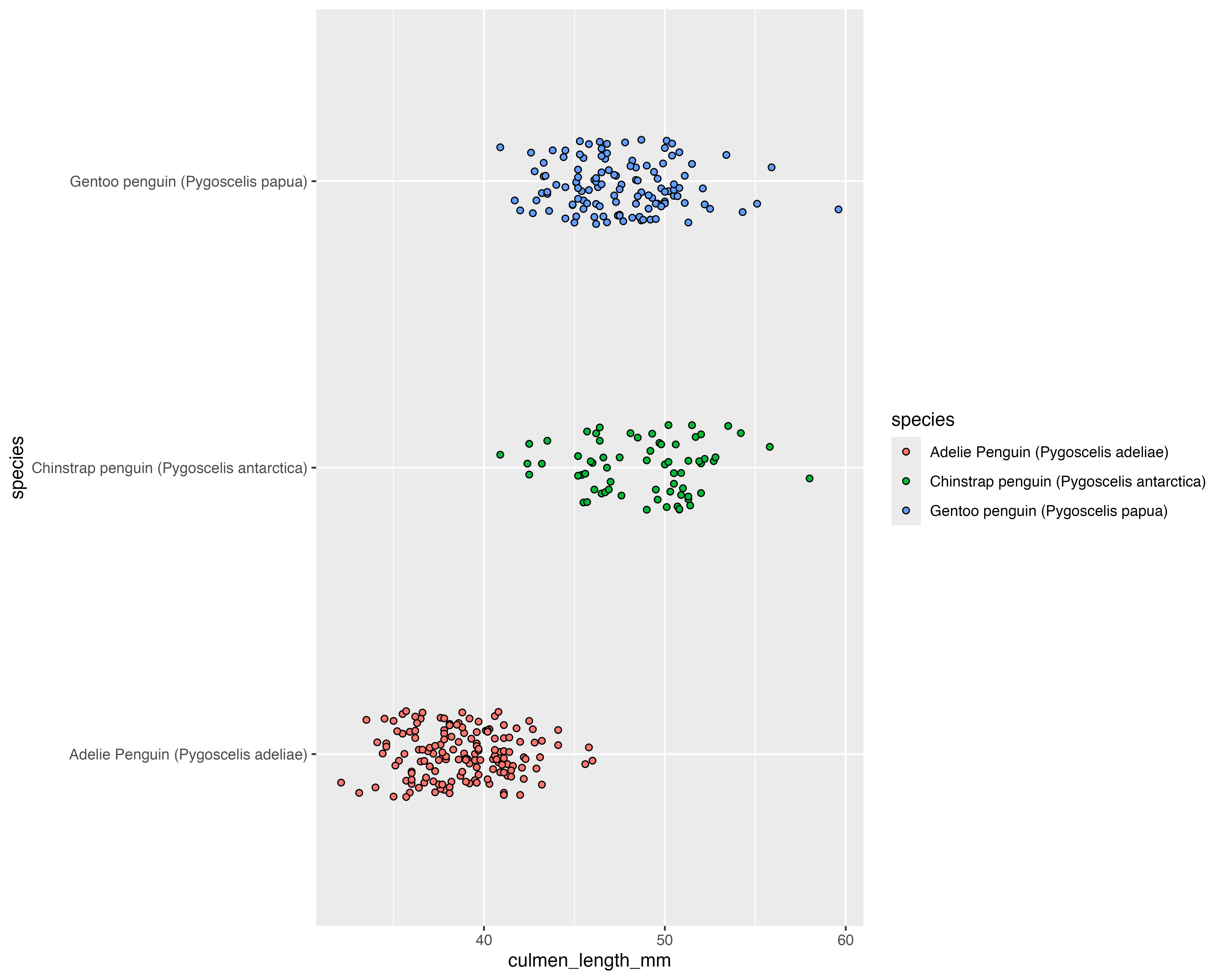
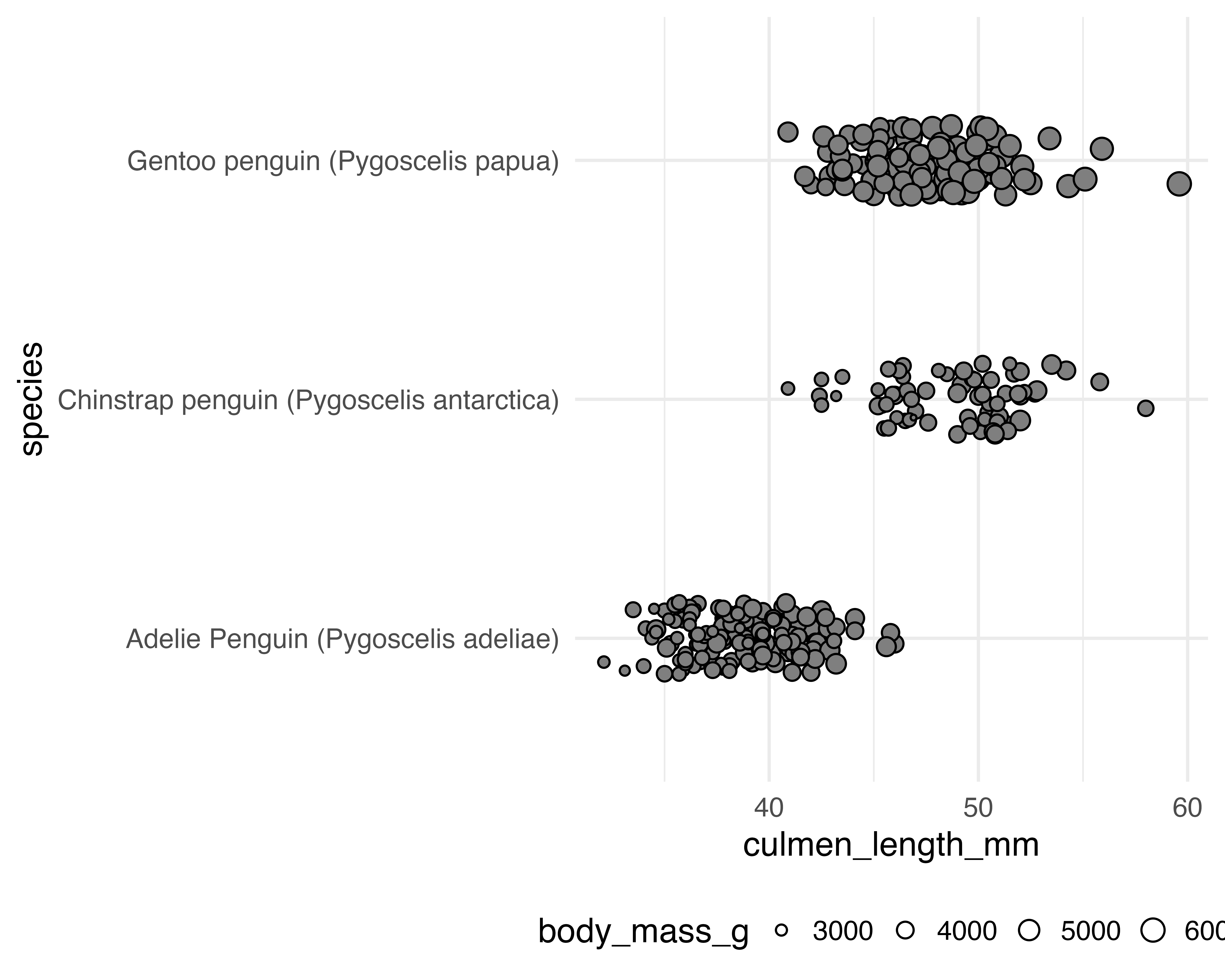
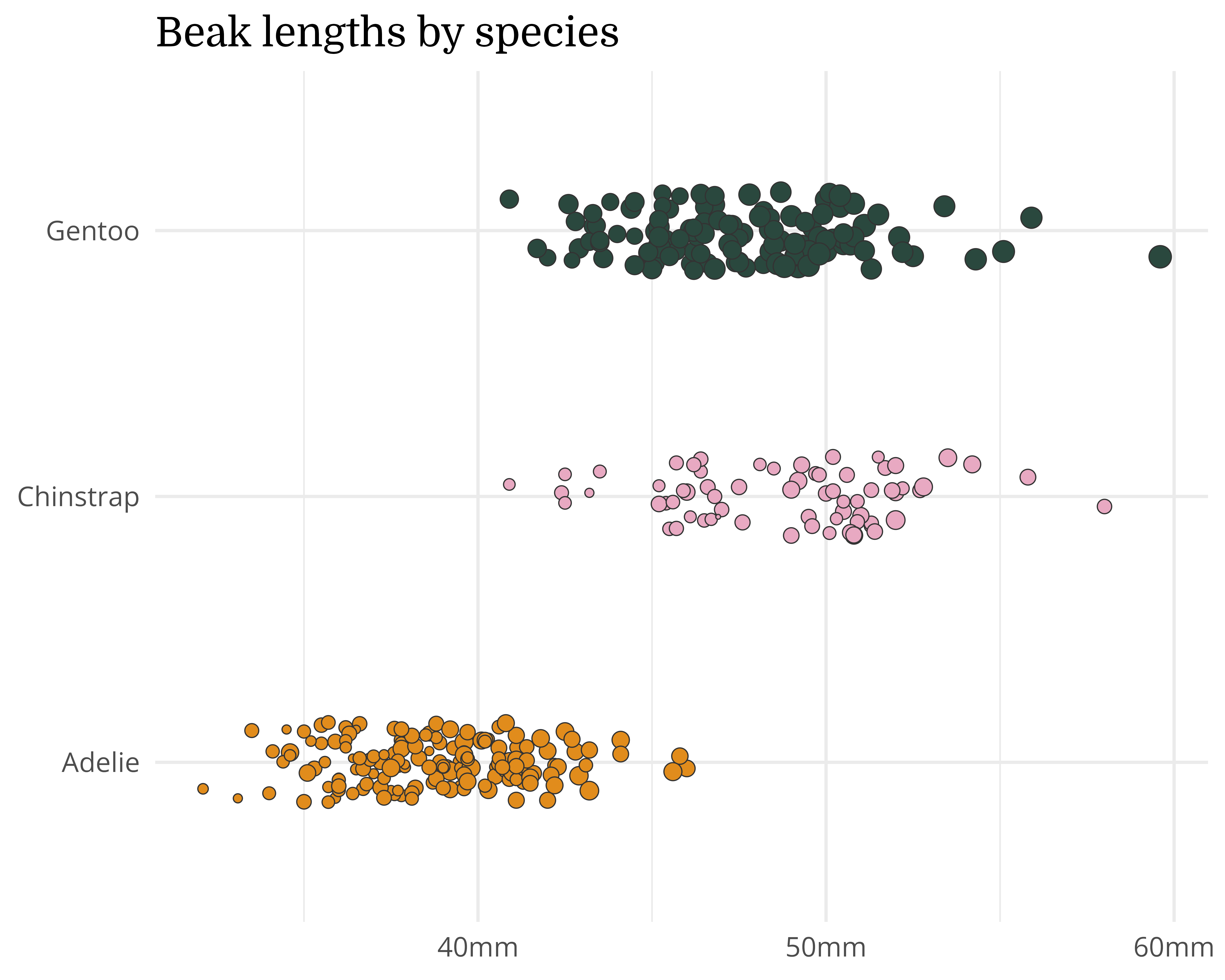
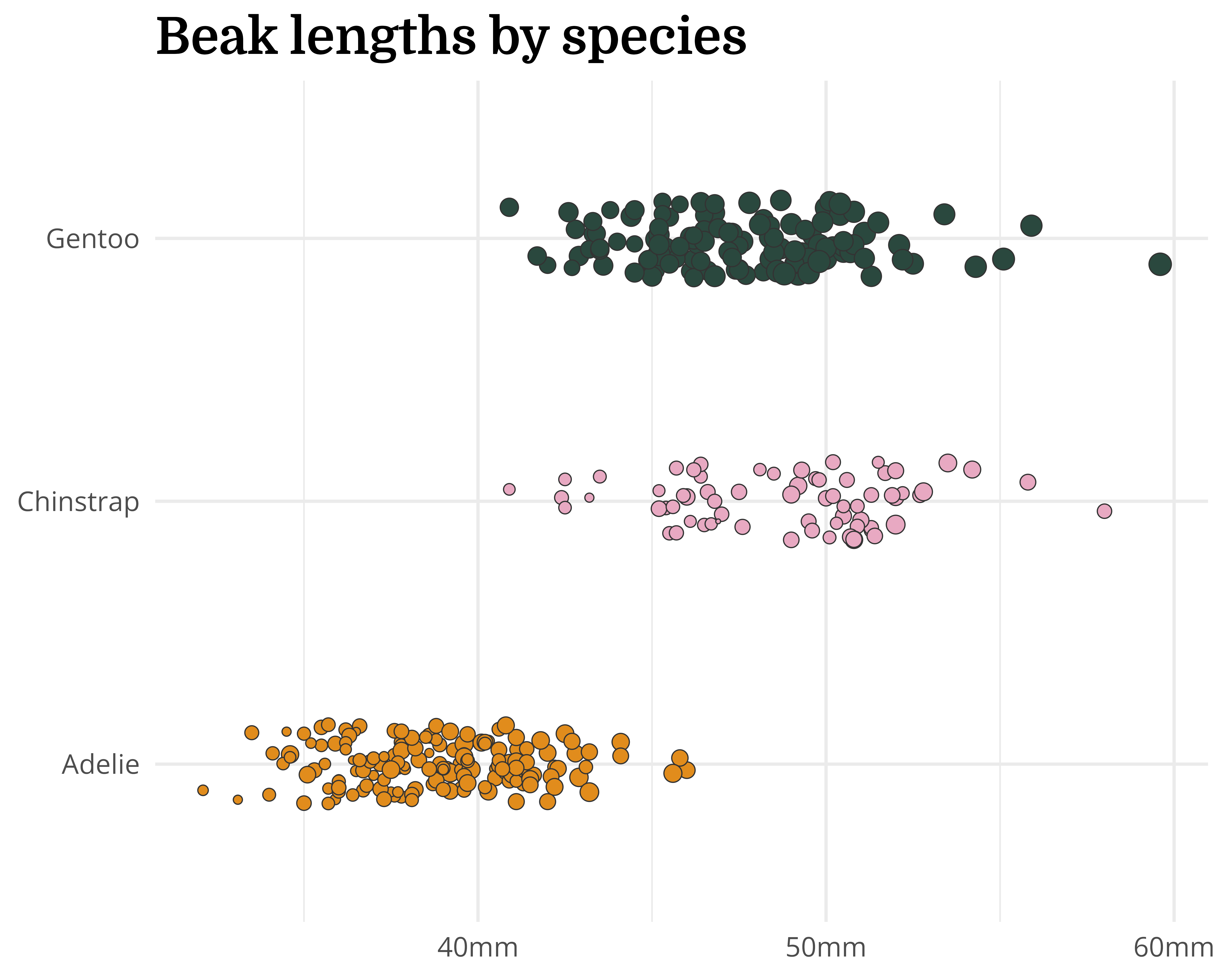
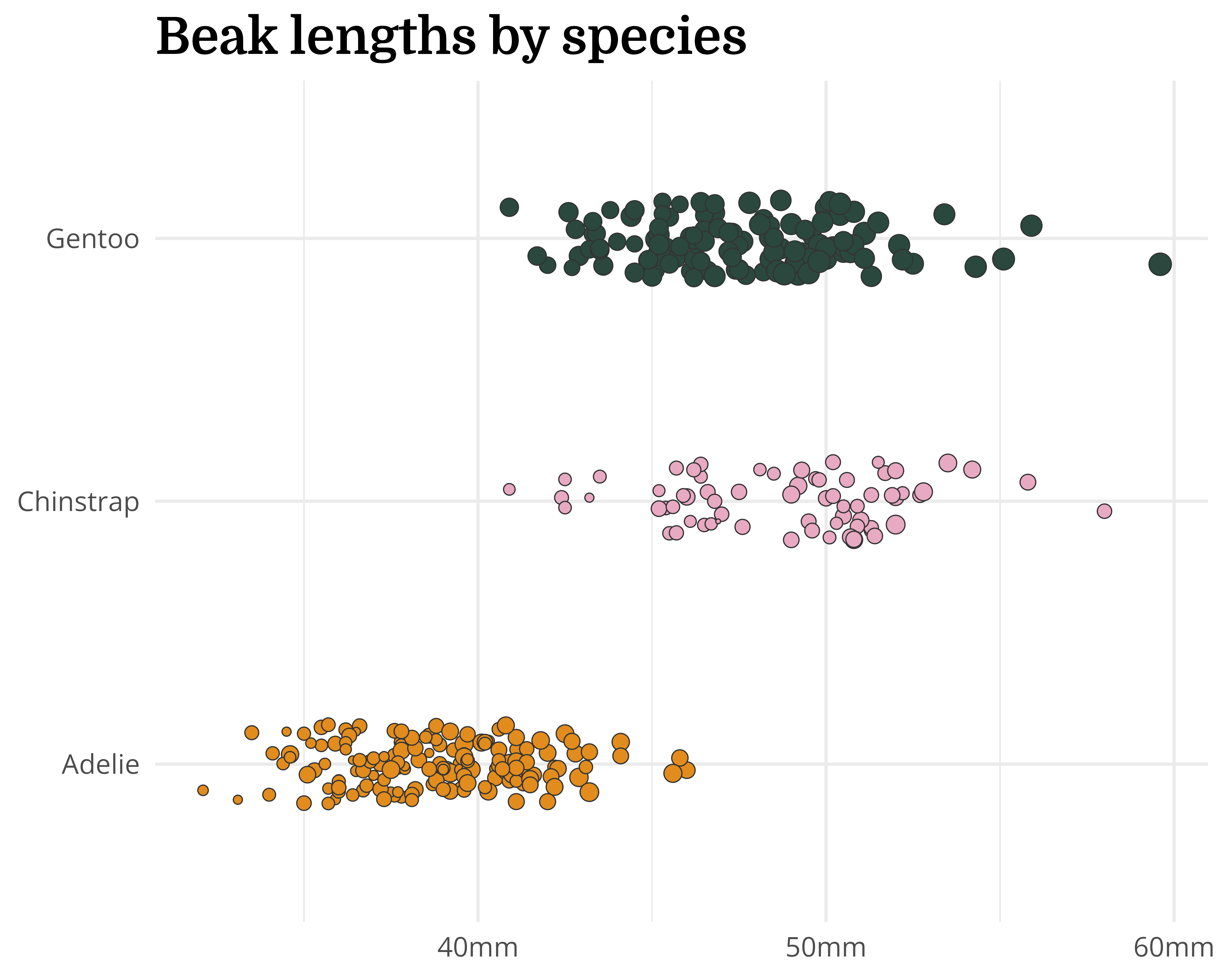
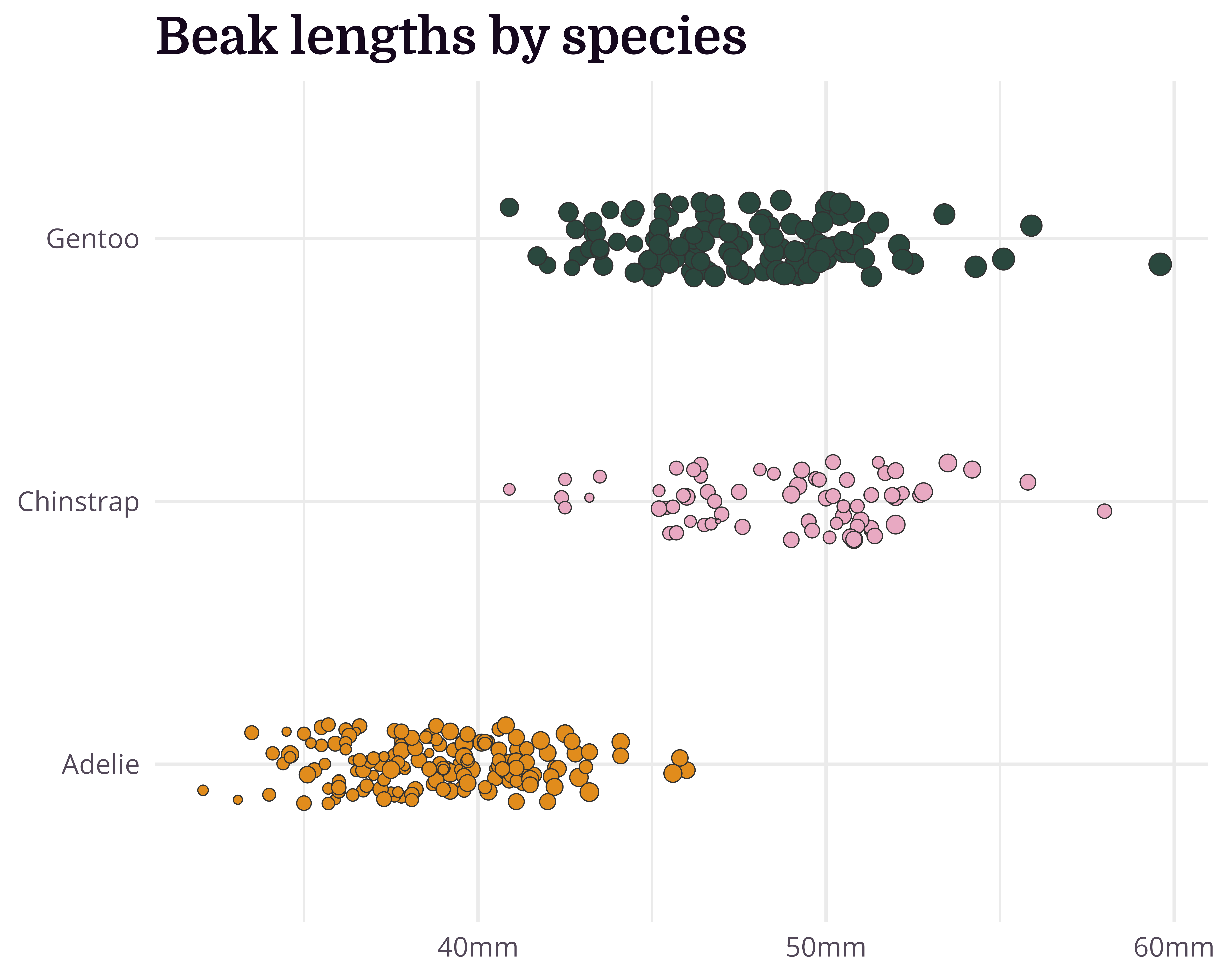
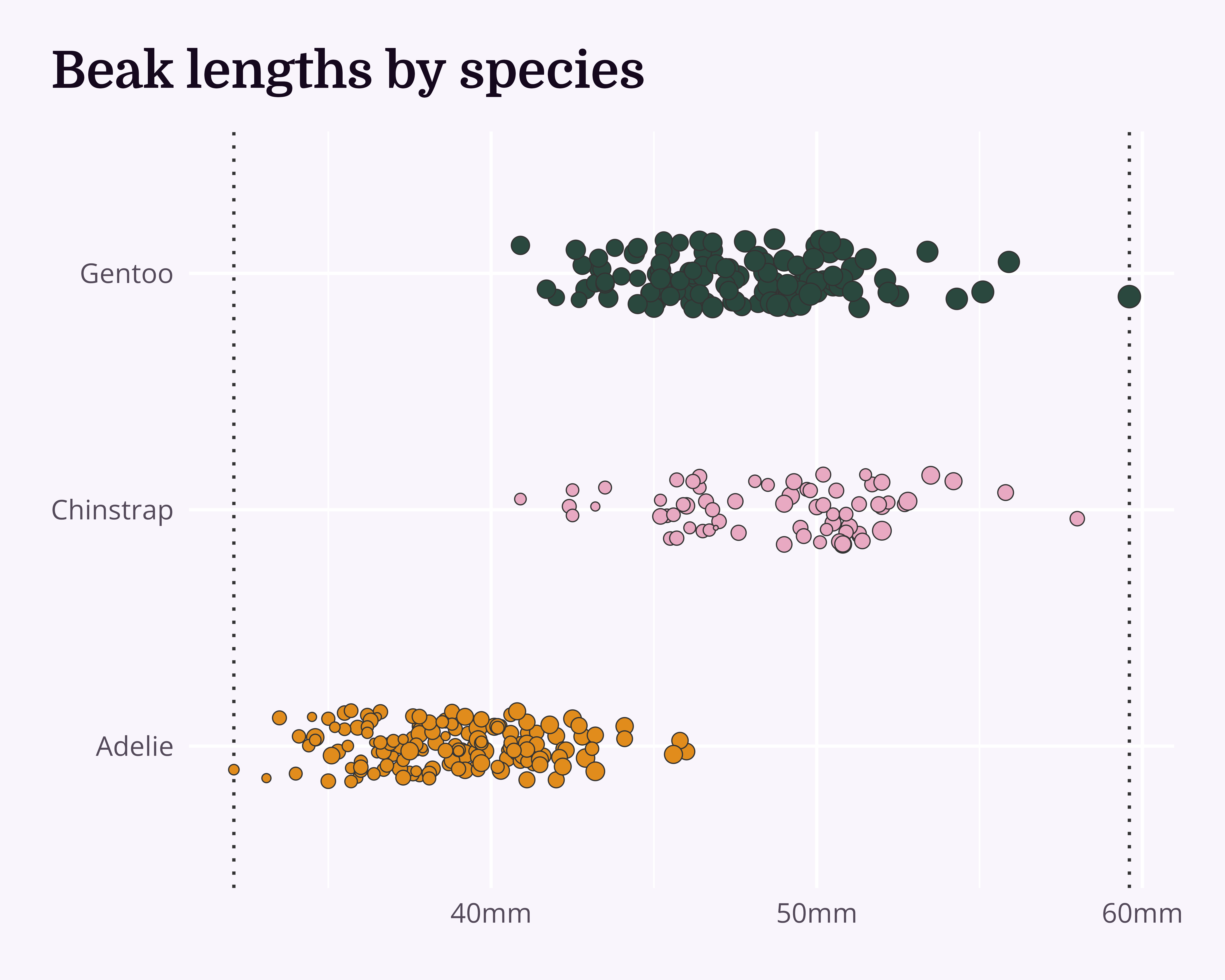
Our starting point
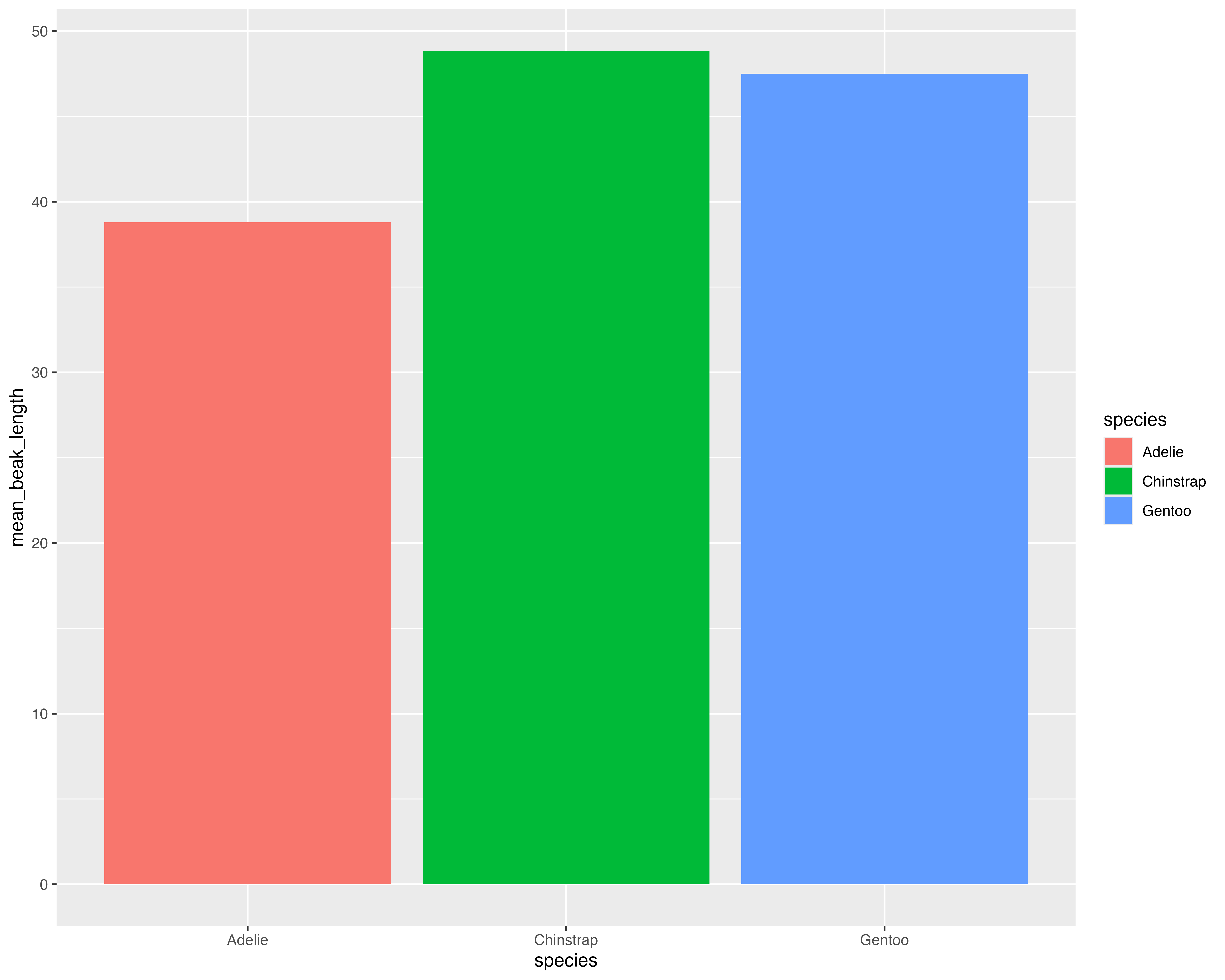
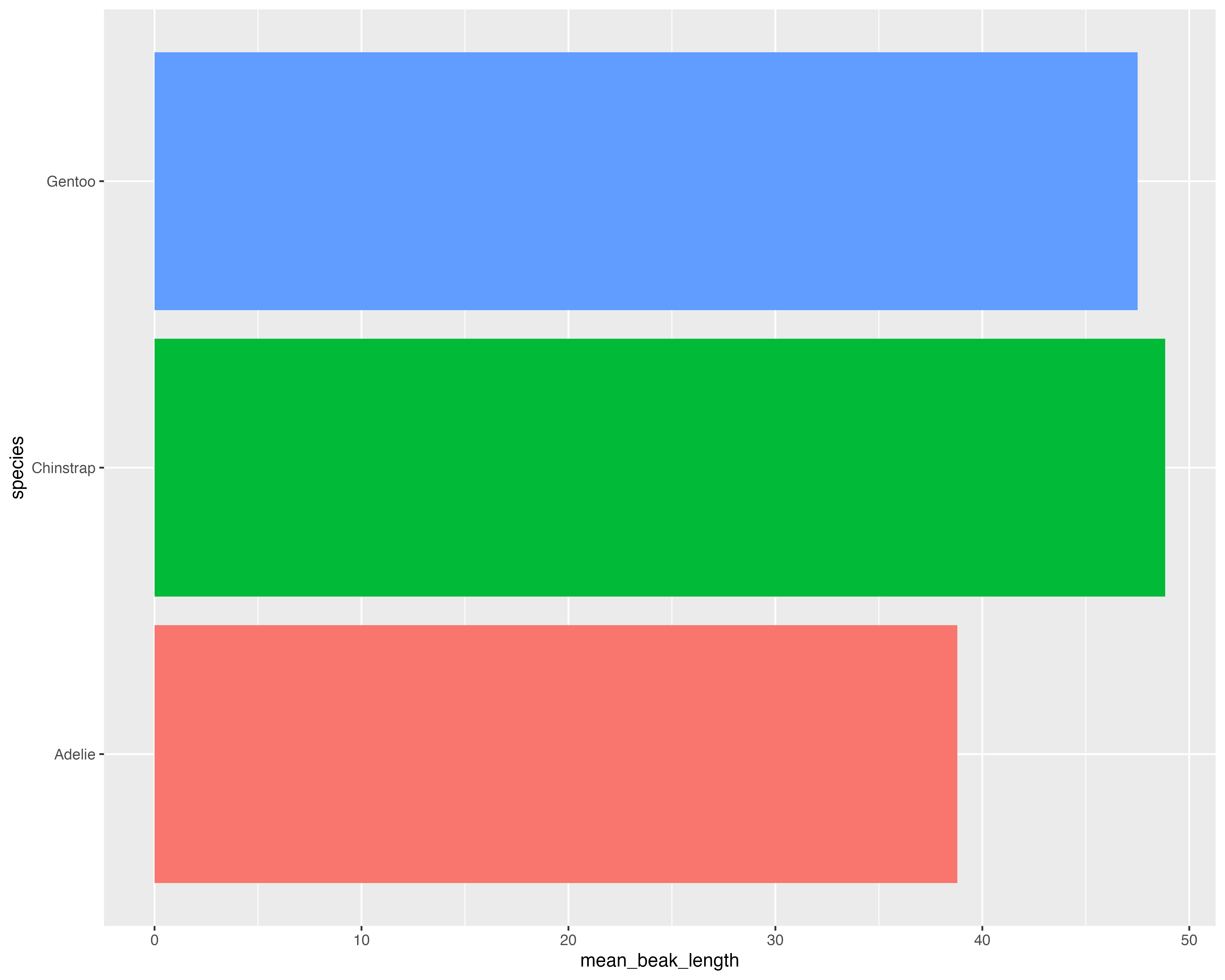
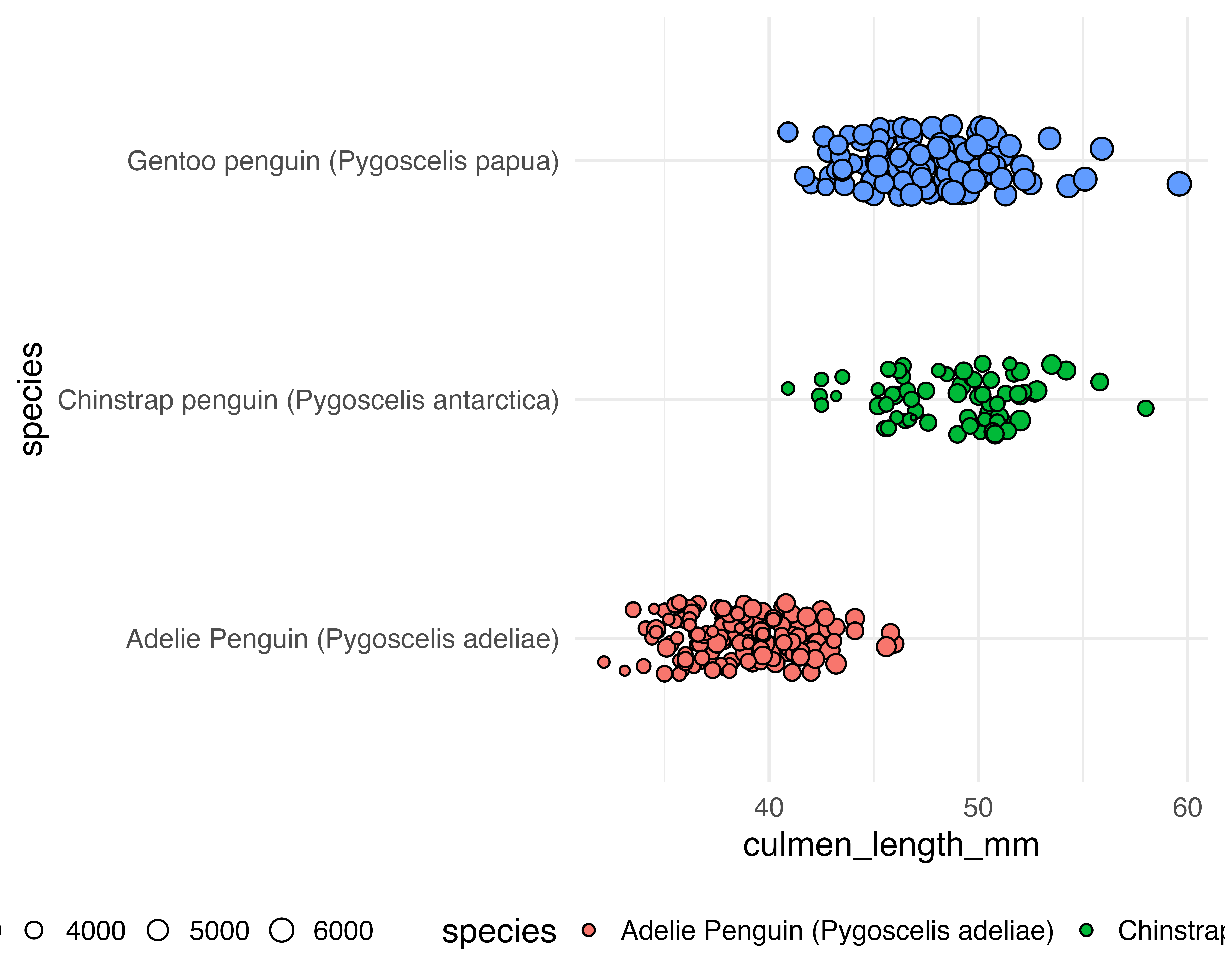
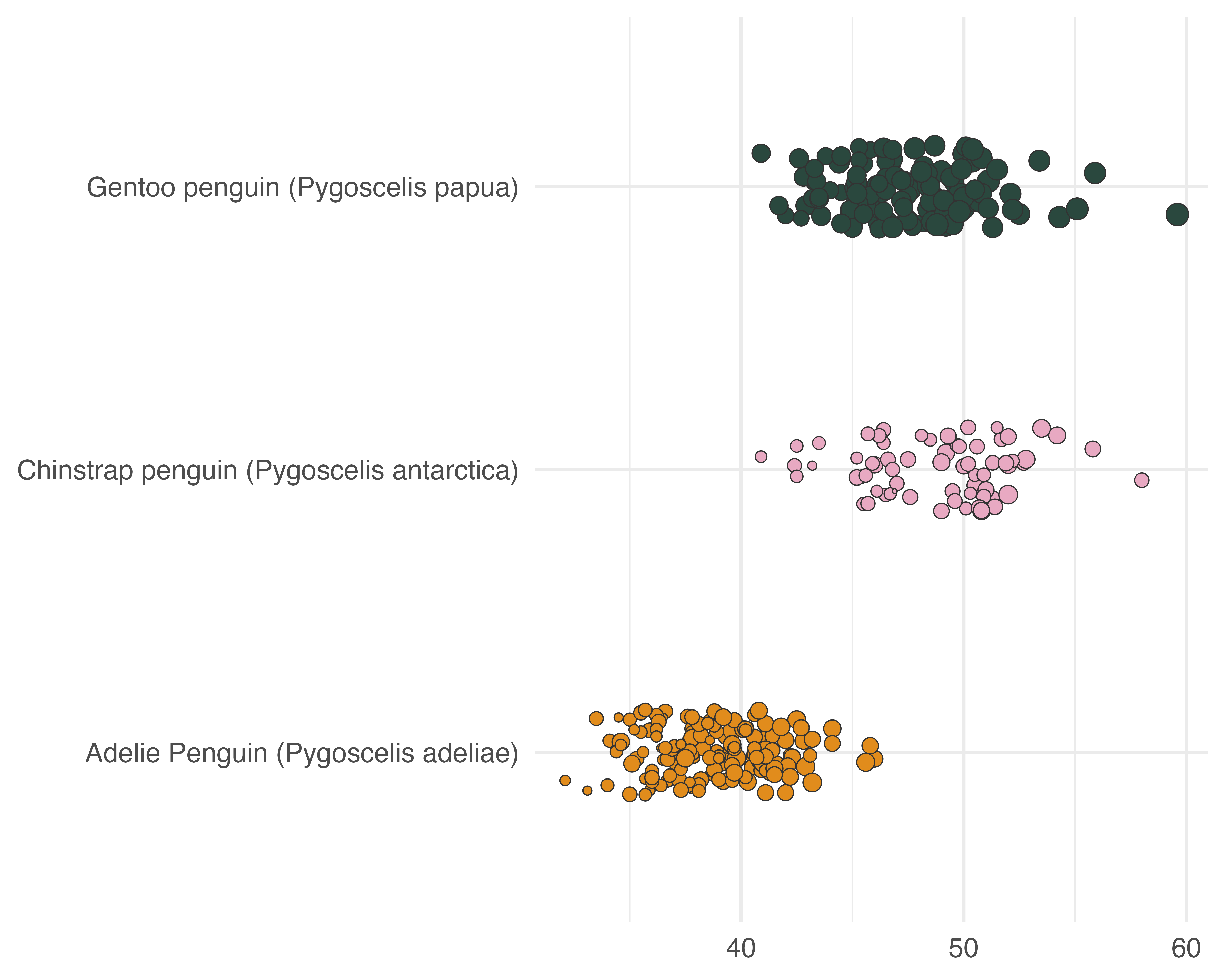
Let’s make a better graph!
Our starting point
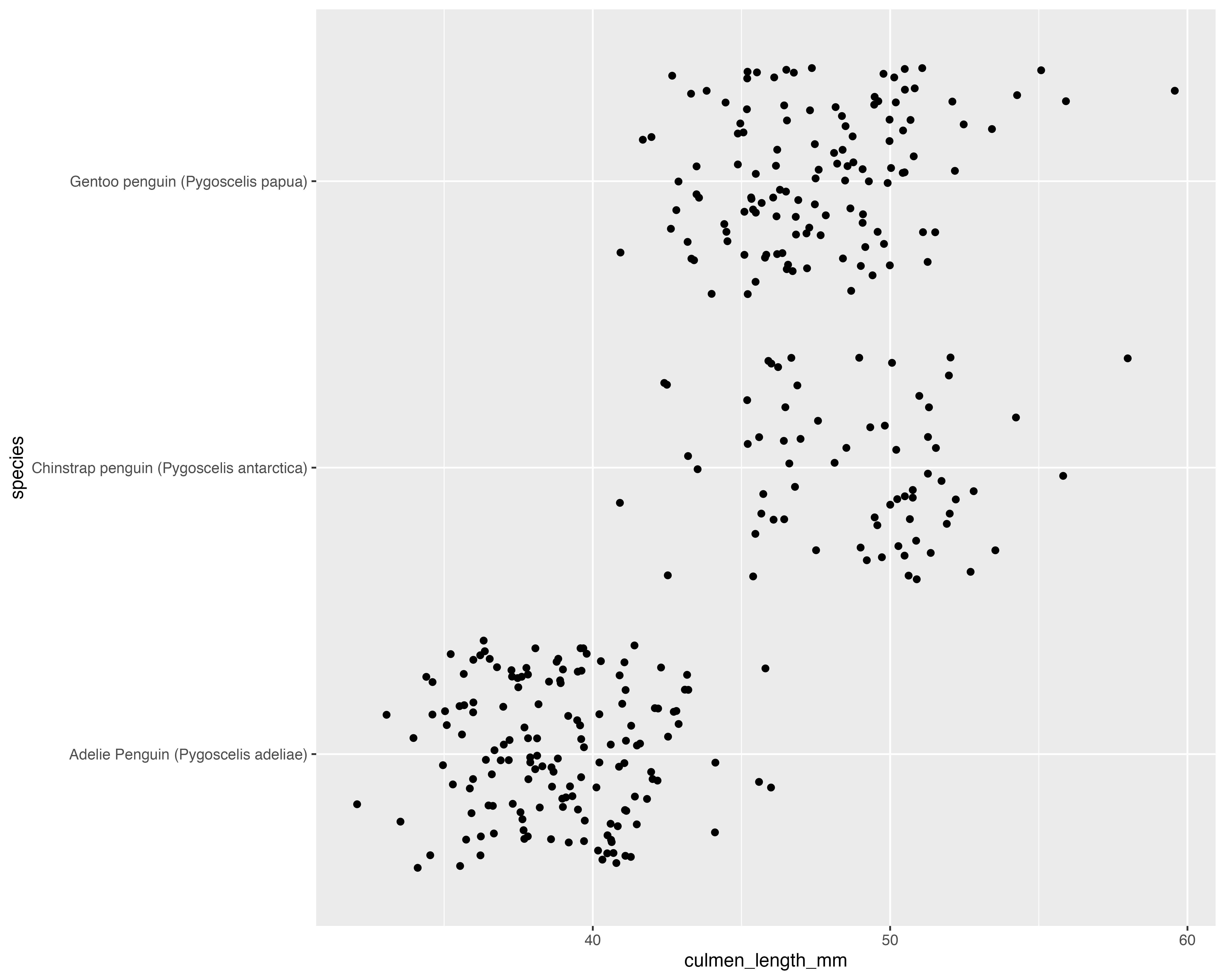
Avoid all the overlaps
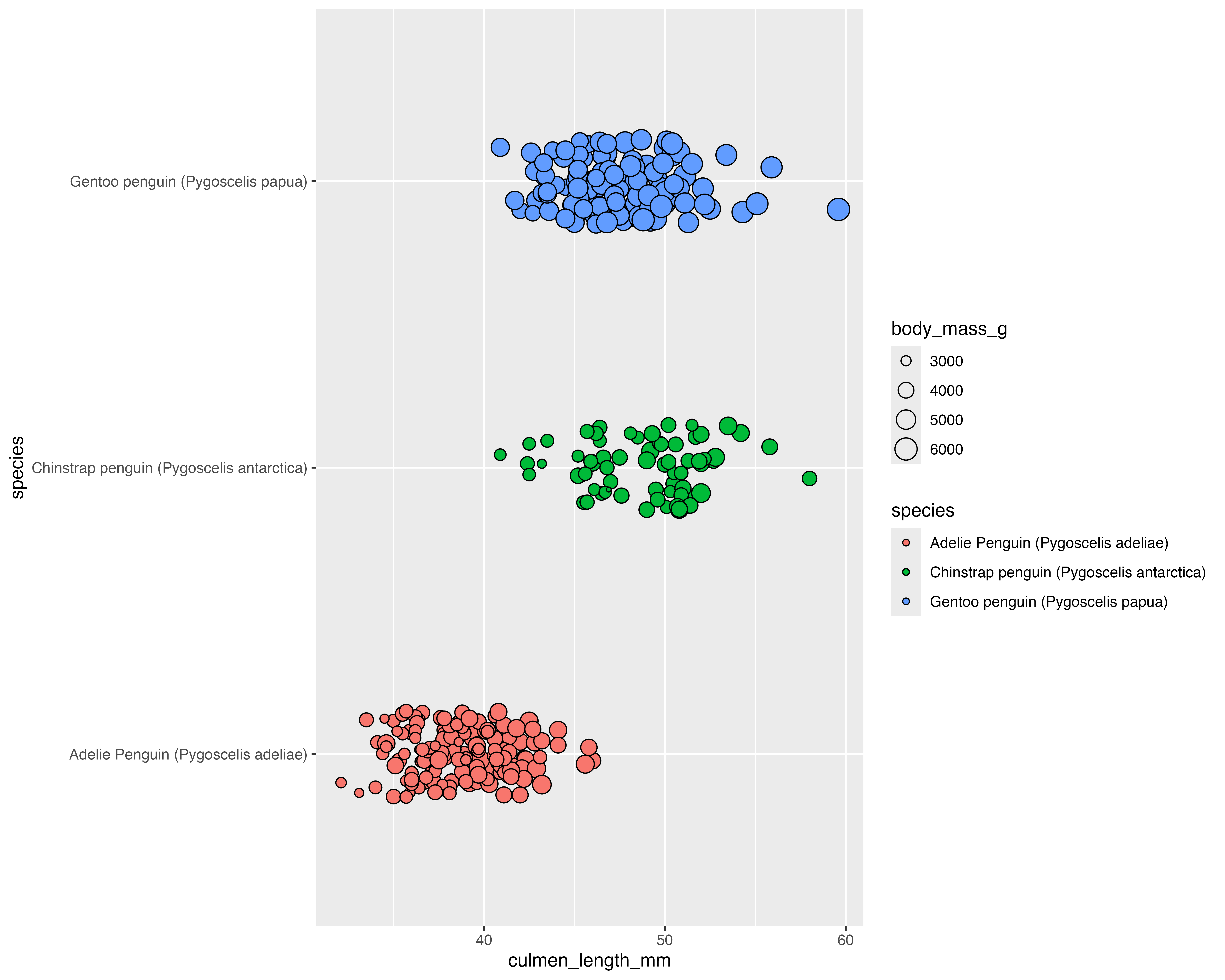
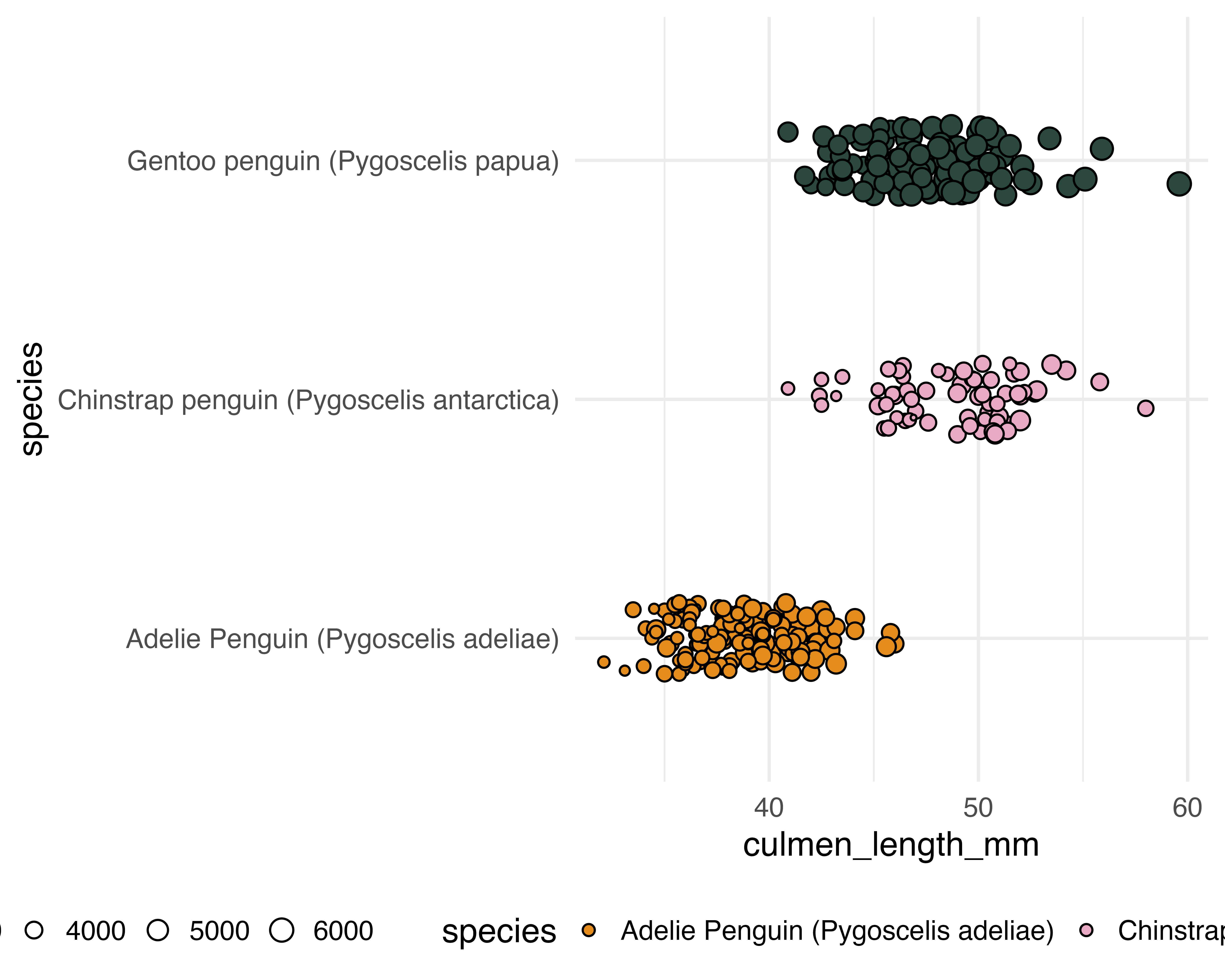
Our starting point
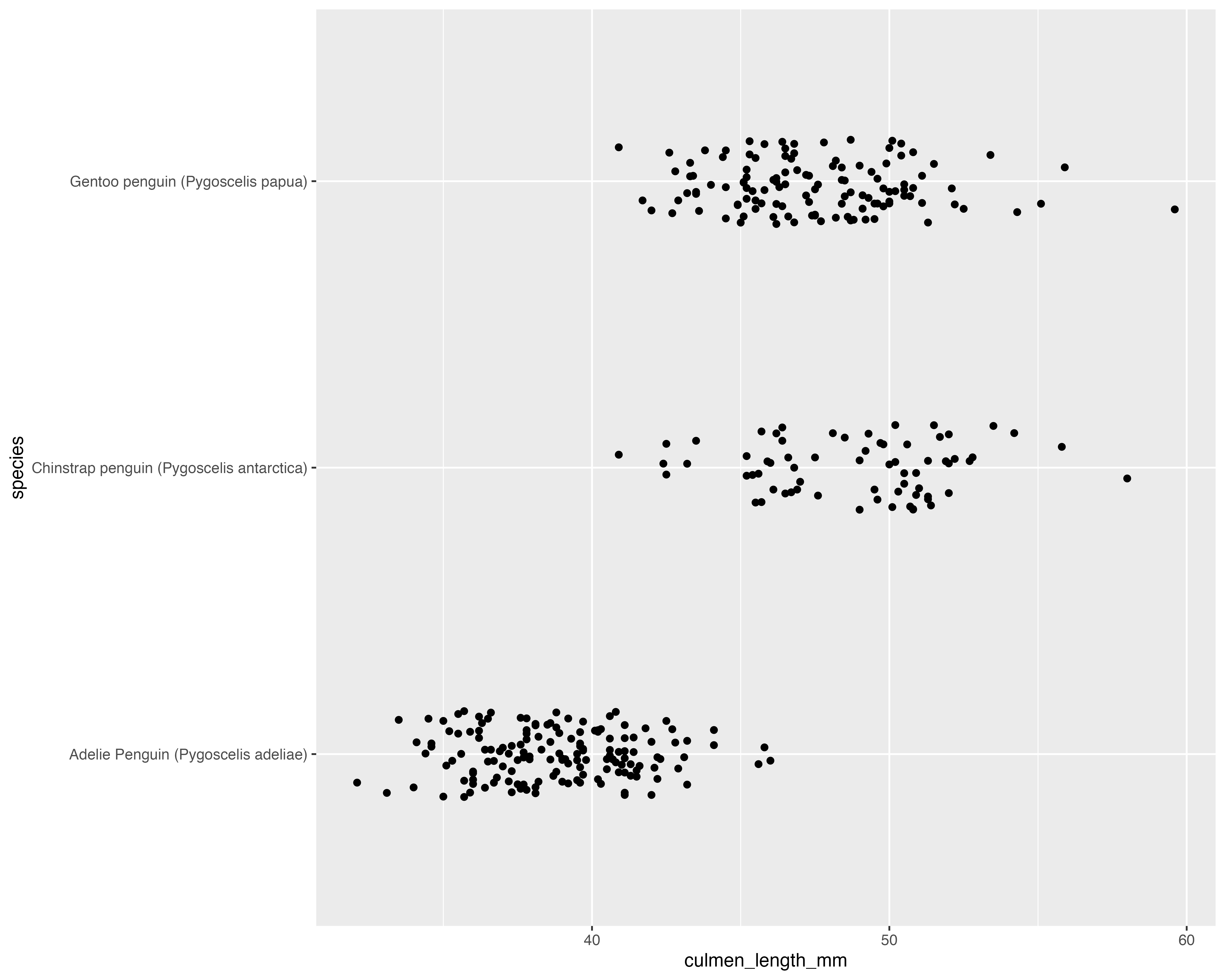
Make the grouping clear, and only jitter what doesn’t matter!
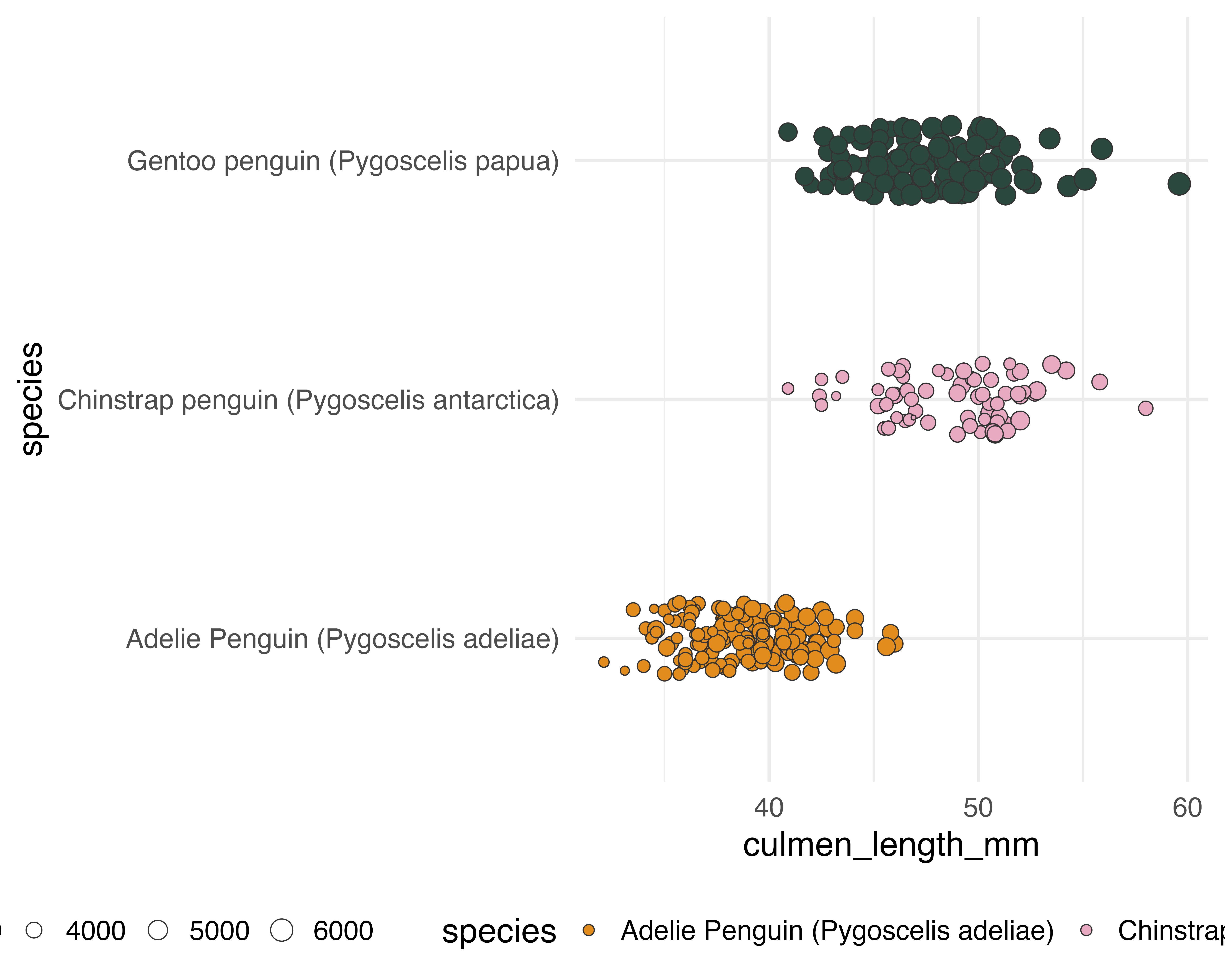
Our starting point
Add a few layers of meaning…
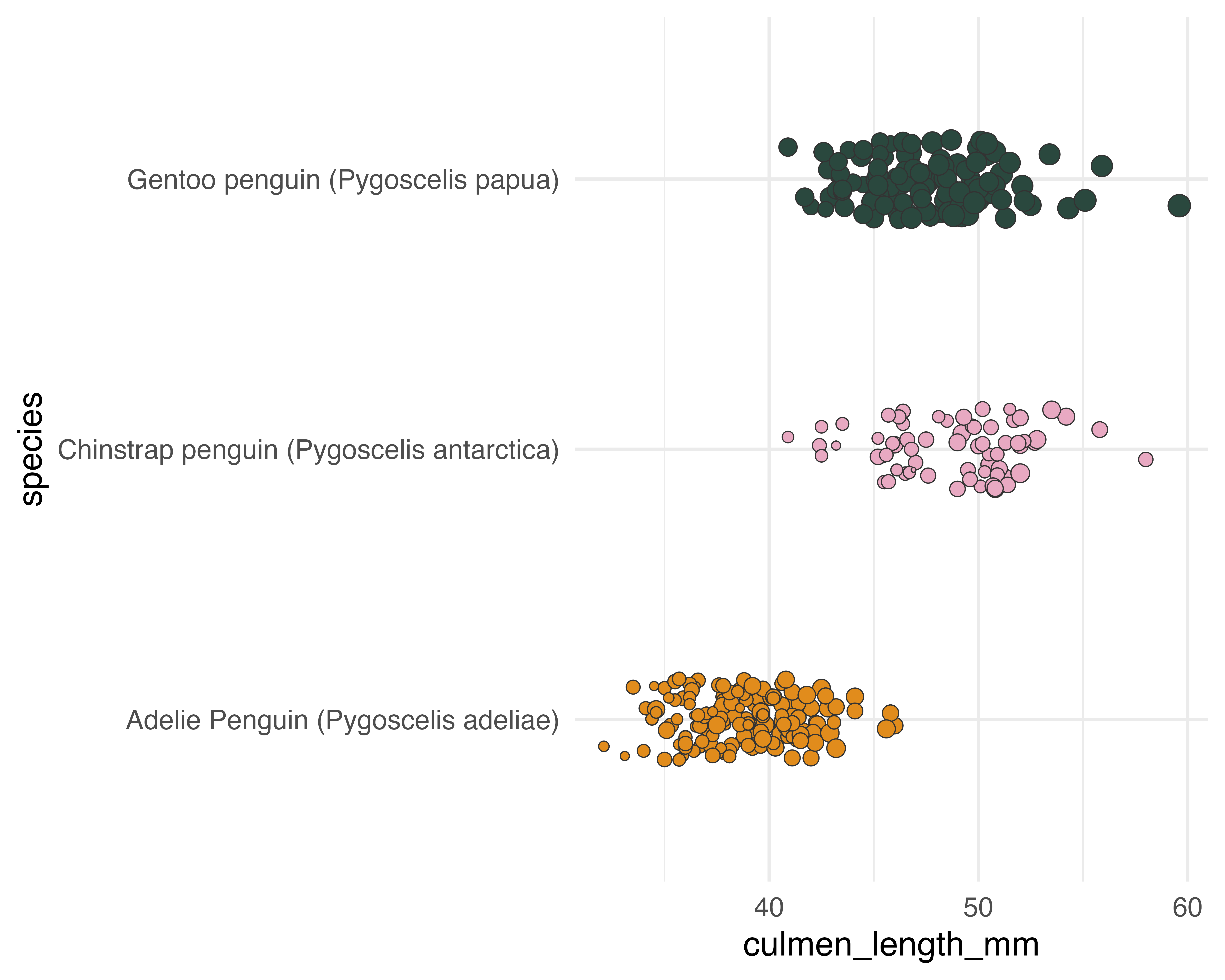
Our starting point
Add a few layers of meaning…
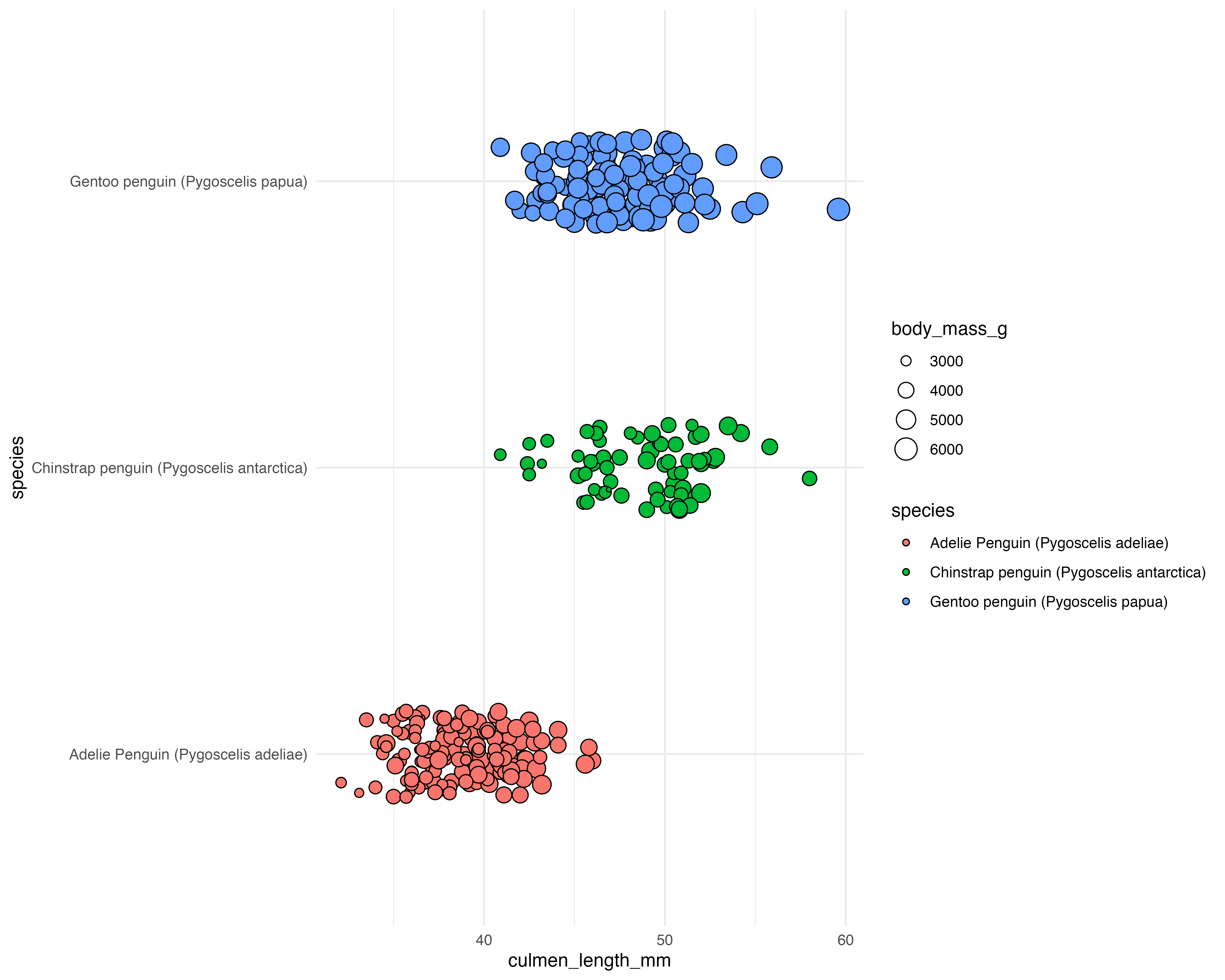
Our starting point
theme_minimal()
Our starting point
theme_minimal()
Our starting point
Move the legend
Optimise!
Colours, text, annotations
Better colours
Better colours
Blending in a common colour
Better colours
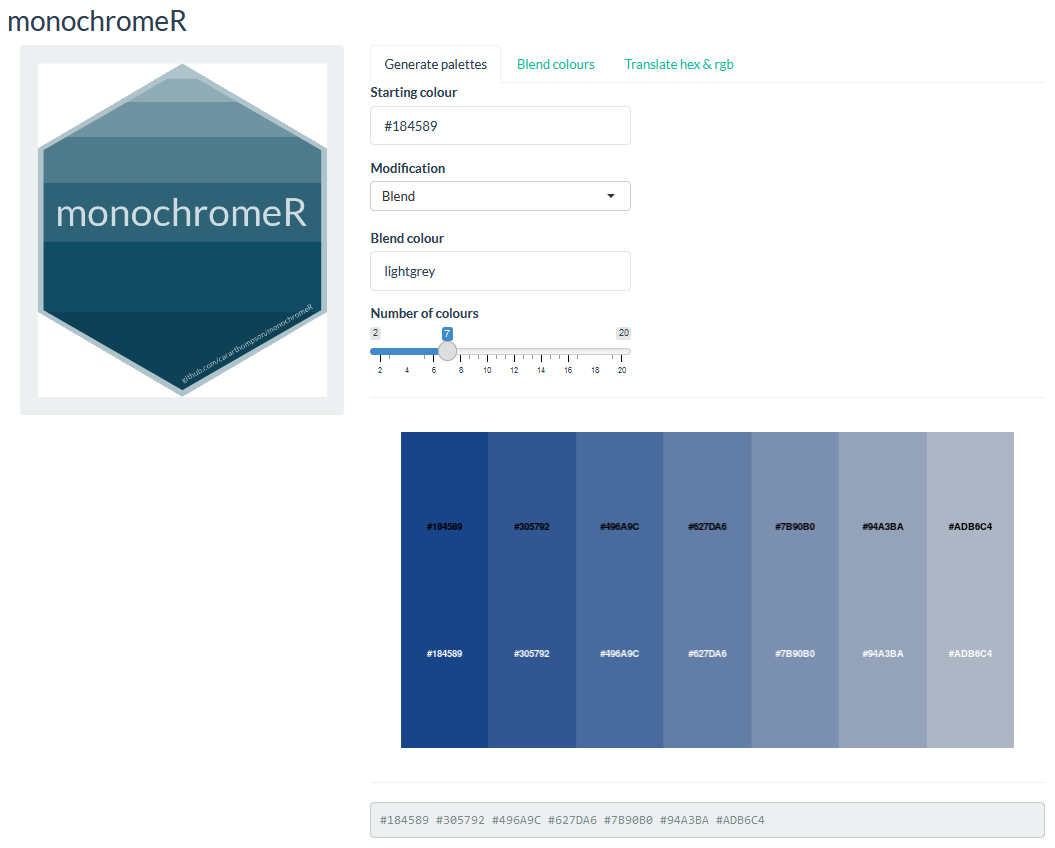
Blending in a common colour - {monochromeR}
Better colours
Blending in a common colour - {monochromeR}
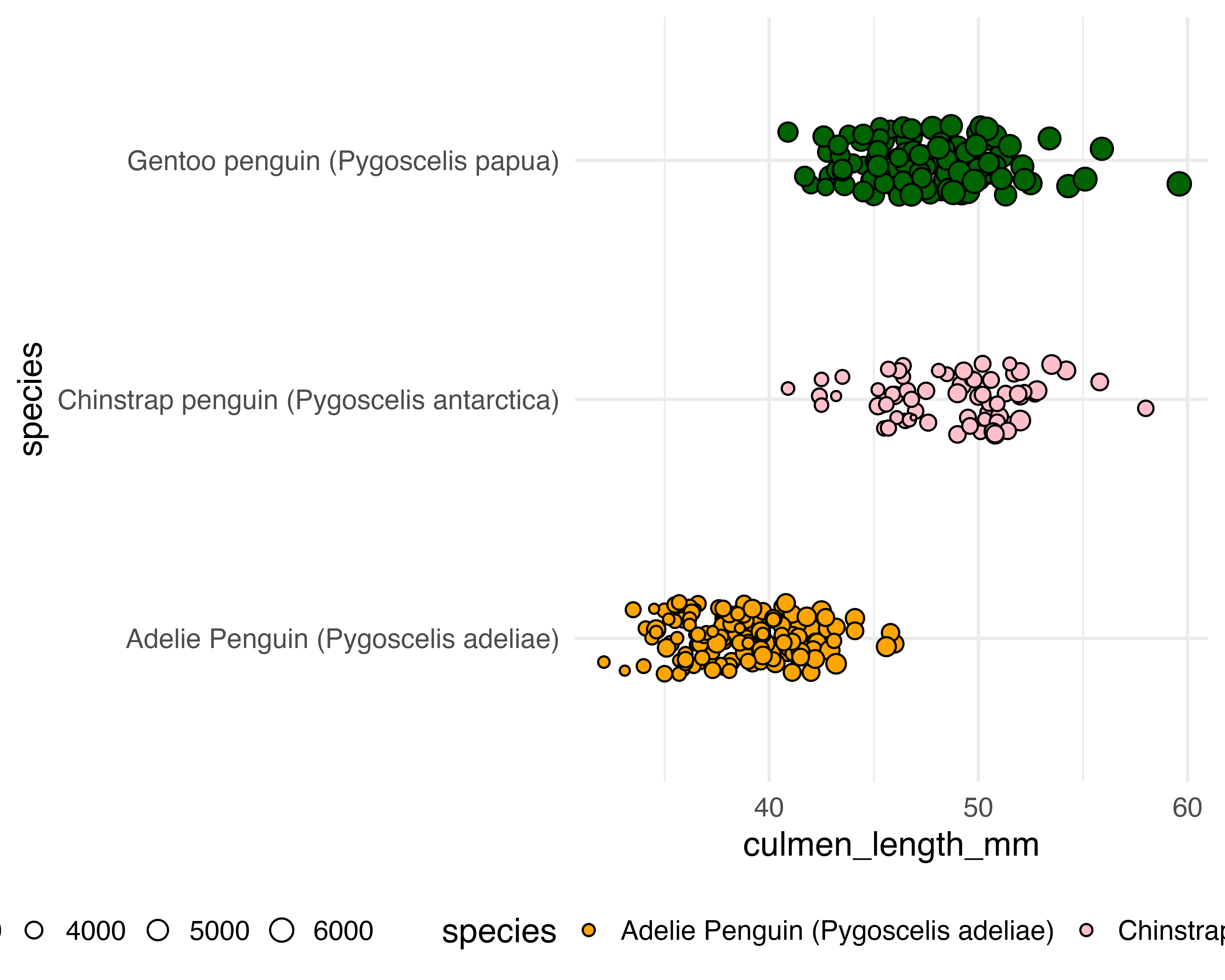
Better colours
It’s subtle… Wait for it!
Better colours
It’s subtle… Wait for it!
Oops!
Named vector needs to match the data!
Better colours
It’s subtle… Wait for it!
Better colours
It’s subtle… One last thing for now
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
scale_fill_manual(values = penguin_colours) +
theme_minimal(base_size = 20) +
theme(legend.position = "bottom")Better text
Did we need the legend? (maybe later?)
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
scale_fill_manual(values = penguin_colours) +
theme_minimal(base_size = 20) +
theme(legend.position = "none")Better text
Or the axis titles?
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
scale_fill_manual(values = penguin_colours) +
theme_minimal(base_size = 20) +
theme(legend.position = "none", axis.title = element_blank())Better text
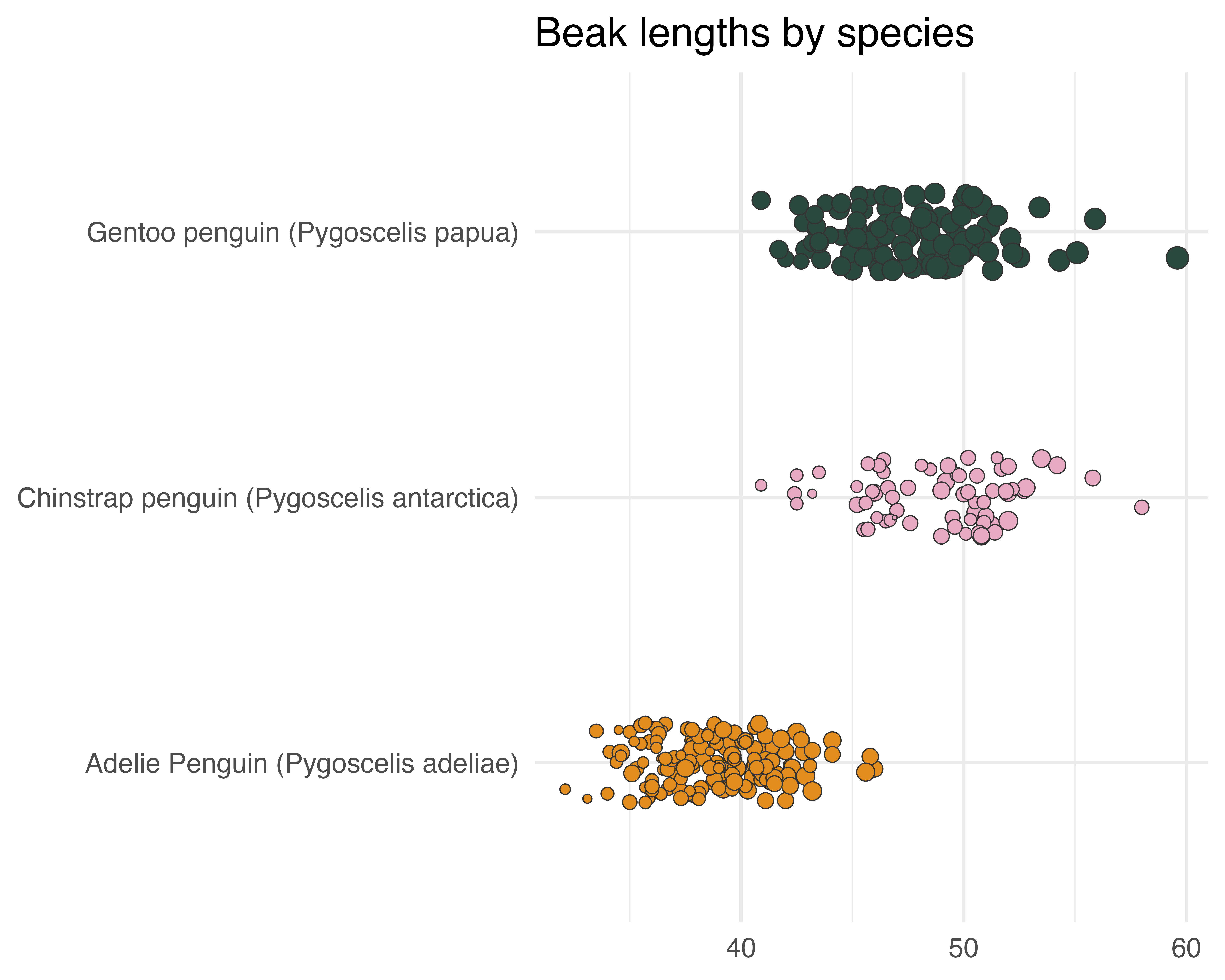
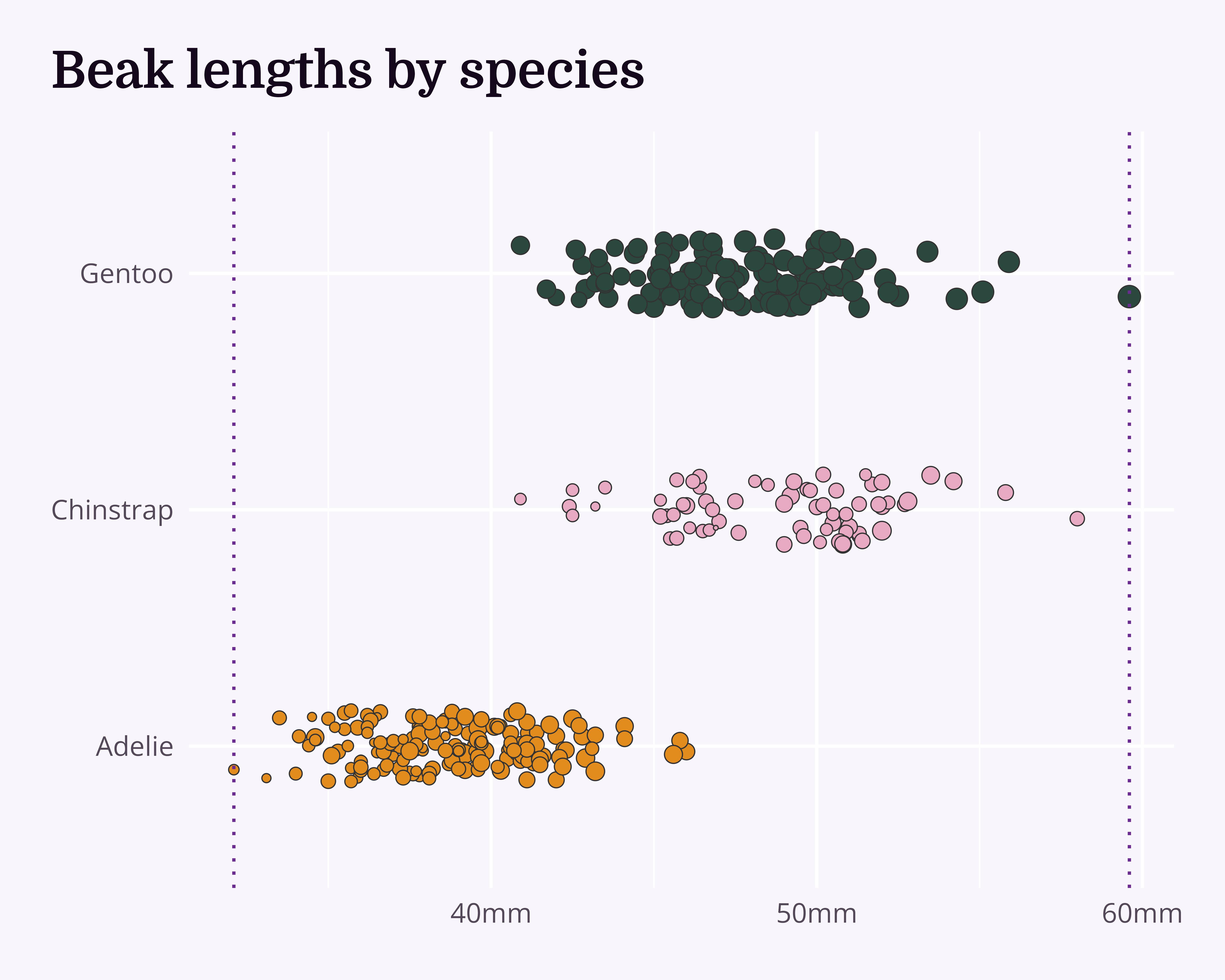
What about a title?
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
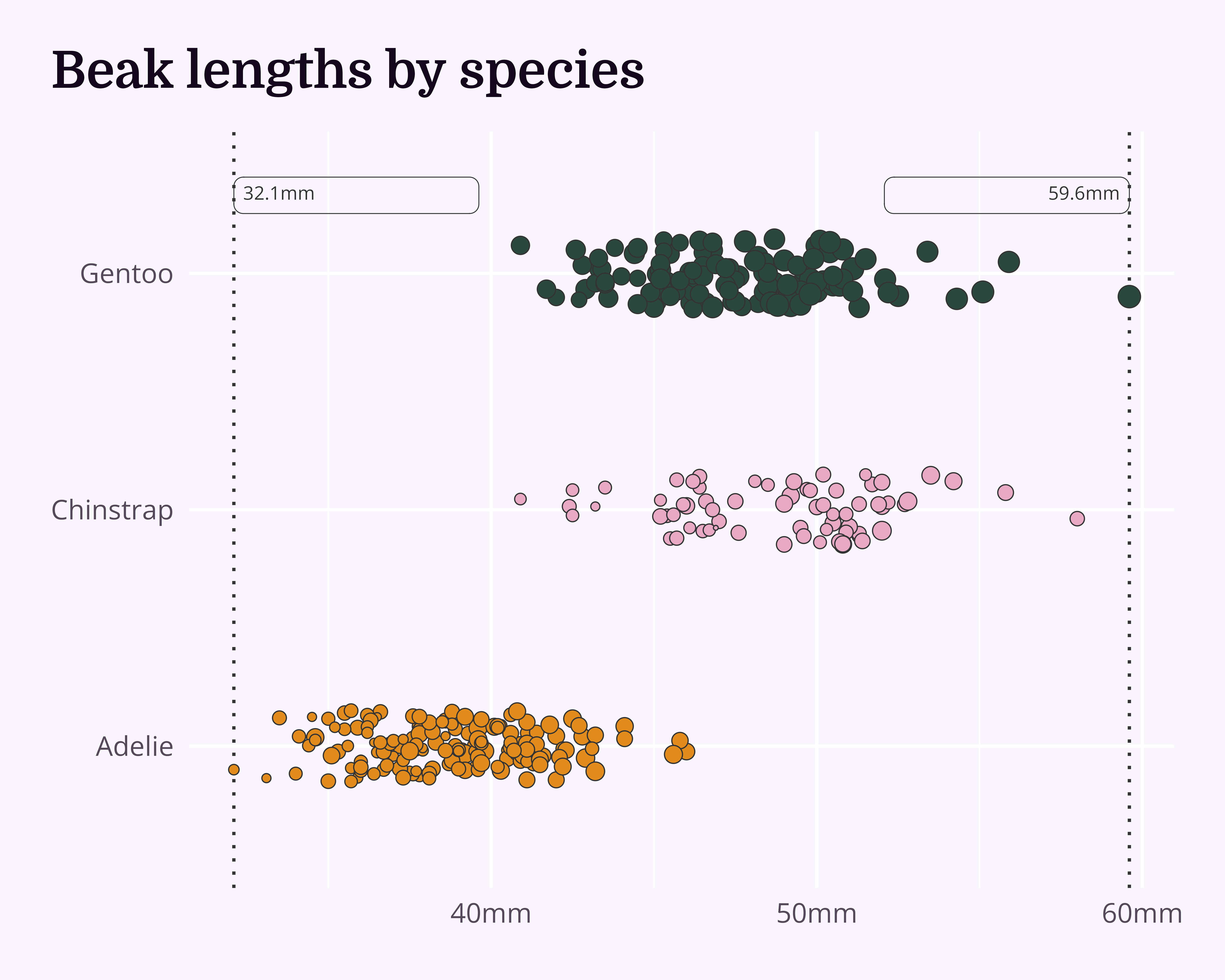
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
theme_minimal(base_size = 20) +
theme(legend.position = "none", axis.title = element_blank())Better text
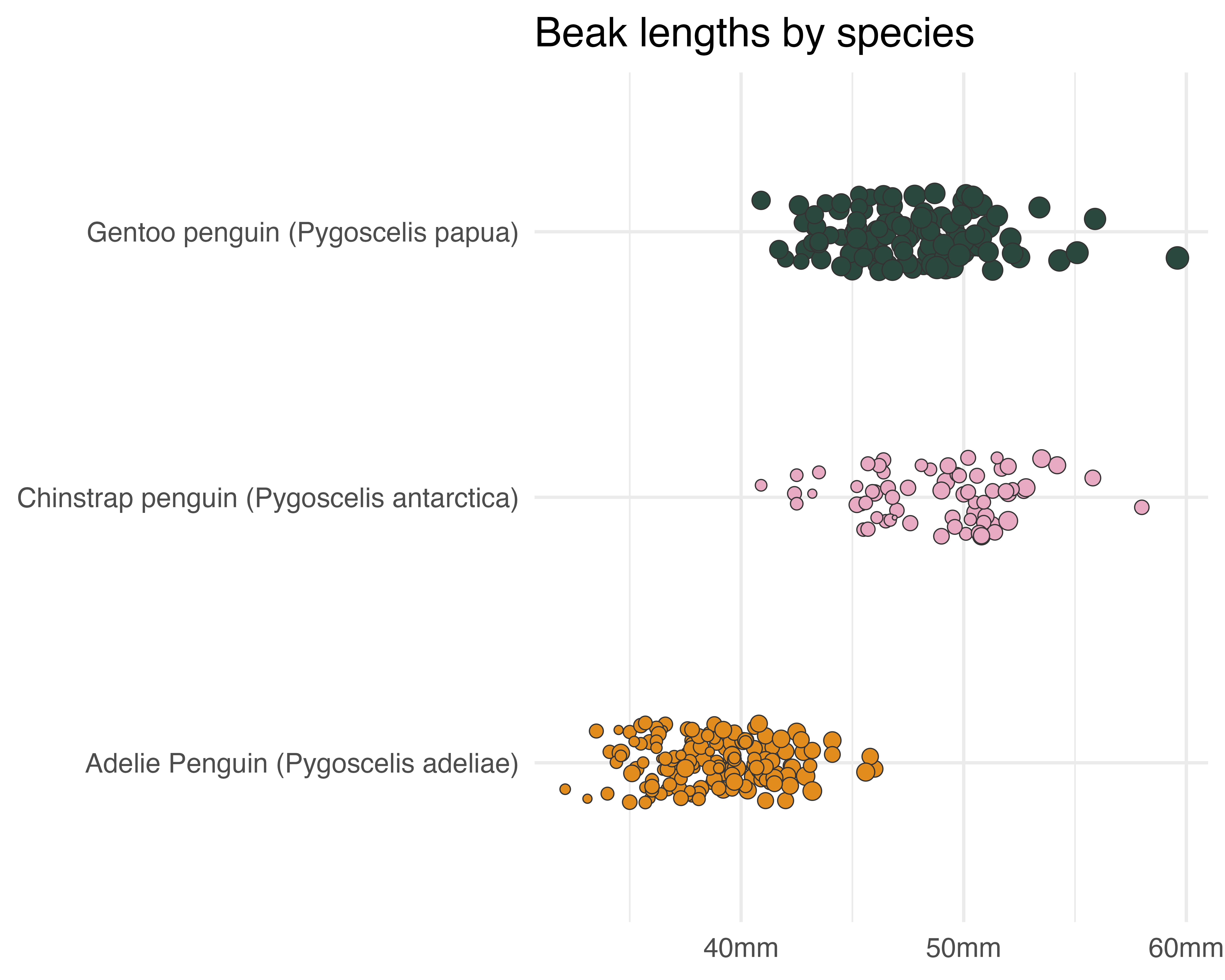
Let’s be helpful
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
theme_minimal(base_size = 20) +
theme(legend.position = "none", axis.title = element_blank())Better text
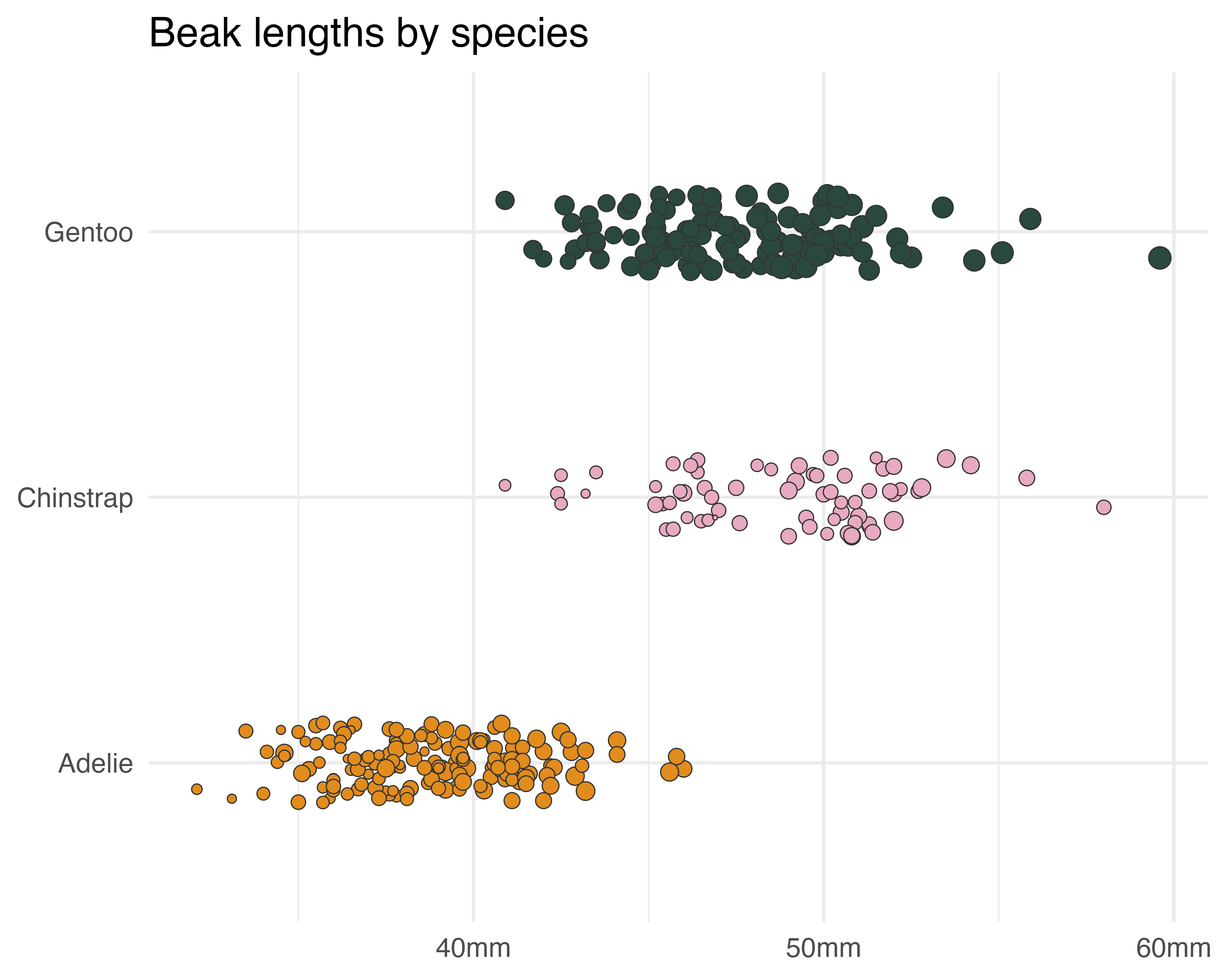
Sort out the y axis text
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_minimal(base_size = 20) +
theme(legend.position = "none", axis.title = element_blank())Better text

Better text
Personality
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_minimal(base_size = 20) +
theme(
text = element_text(family = "Open Sans"),
legend.position = "none",
axis.title = element_blank(),
plot.title = element_text(family = "Domine")
)Better text
Personality + hierarchy
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_minimal(base_size = 20) +
theme(
text = element_text(family = "Open Sans"),
legend.position = "none",
axis.title = element_blank(),
plot.title = element_text(
family = "Domine",
face = "bold",
size = 30
)
)Better text
Personality + hierarchy (better!)
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_minimal(base_size = 20) +
theme(
text = element_text(family = "Open Sans"),
legend.position = "none",
axis.title = element_blank(),
plot.title = element_text(
family = "Domine",
face = "bold",
size = rel(1.5)
)
)Better text
Personality + hierarchy + colour
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_minimal(base_size = 20) +
theme(
text = element_text(family = "Open Sans", colour = "#534959"),
axis.text = element_text(colour = "#534959"),
legend.position = "none",
axis.title = element_blank(),
plot.title = element_text(
family = "Domine",
face = "bold",
size = rel(1.5),
colour = "#15081D"
)
)Getting fonts to work in R
Getting fonts to work can be frustrating!
Install fonts locally, restart R Studio + 📦
{systemfonts}({ragg}+{textshaping}) + Set graphics device to “AGG” + 🤞
knitr::opts_chunk$set(dev = “ragg_png”)
Optimise the small things
Background, grid, margins…
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_minimal(base_size = 20) +
theme(
text = element_text(family = "Open Sans", colour = "#534959"),
axis.text = element_text(colour = "#534959"),
legend.position = "none",
axis.title = element_blank(),
plot.title.position = "plot",
plot.title = element_text(
family = "Domine",
face = "bold",
size = rel(1.5),
colour = "#15081D",
margin = margin(0, 0, 20, 0)
),
panel.grid = element_line(colour = "#FFFFFF"),
plot.background = element_rect(fill = "#f9f5fc", colour = "#f9f5fc"),
plot.margin = margin_auto(30)
)Helping your future self
We can turn all that styling into a theme function!
plot +
theme_minimal(base_size = 20) +
theme(
text = element_text(family = "Open Sans", colour = "#534959"),
axis.text = element_text(colour = "#534959"),
legend.position = "none",
axis.title = element_blank(),
plot.title.position = "plot",
plot.title = element_text(
family = "Domine",
face = "bold",
size = rel(1.5),
colour = "#15081D",
margin = margin(0, 0, 20, 0)
),
panel.grid = element_line(colour = "#FFFFFF"),
plot.background = element_rect(fill = "#f9f5fc", colour = "#f9f5fc"),
plot.margin = margin_auto(30)
)Helping your future self
We can turn all that styling into a theme function!
theme_chester_penguins <- function(base_text_size = 20) {
theme_minimal(base_size = base_text_size) +
theme(
text = element_text(family = "Open Sans", colour = "#534959"),
axis.text = element_text(colour = "#534959"),
legend.position = "none",
axis.title = element_blank(),
plot.title.position = "plot",
plot.title = ggtext::element_textbox_simple(
family = "Domine",
face = "bold",
size = rel(1.5),
colour = "#15081D",
margin = margin(0, 0, base_text_size, 0)
),
panel.grid = element_line(colour = "#FFFFFF"),
plot.background = element_rect(fill = "#f9f5fc", colour = "#f9f5fc"),
plot.margin = margin_auto(base_text_size * 1.5),
geom = element_geom(ink = "#6b2c91")
)
}Helping your future self
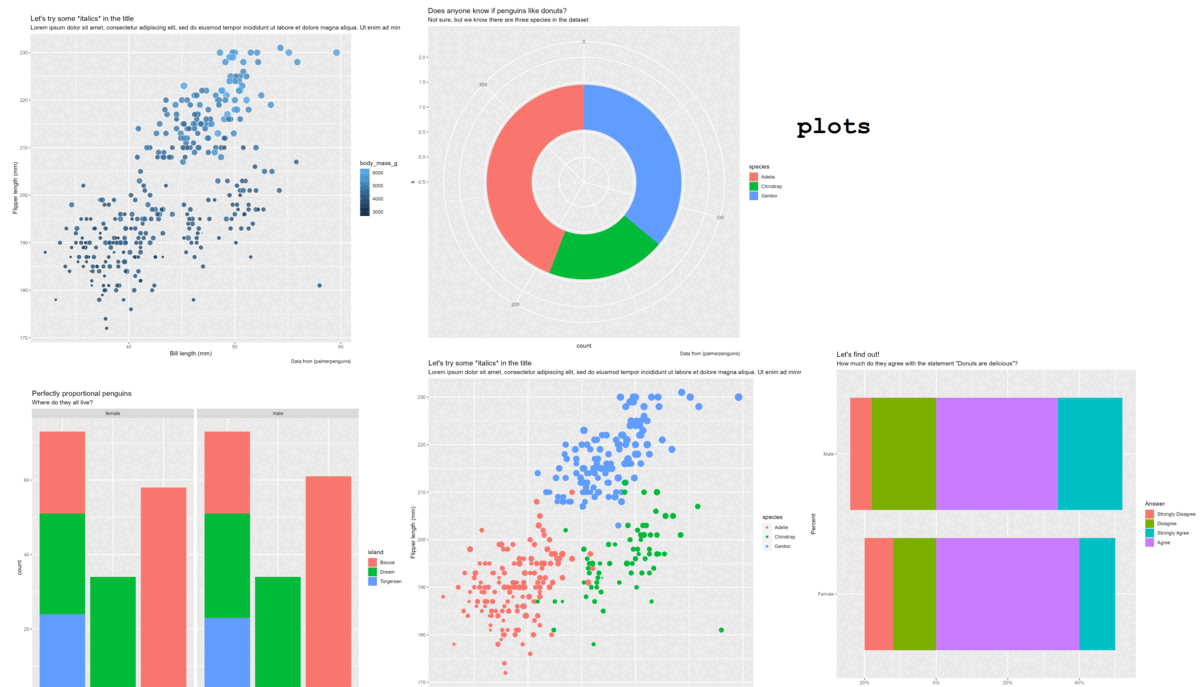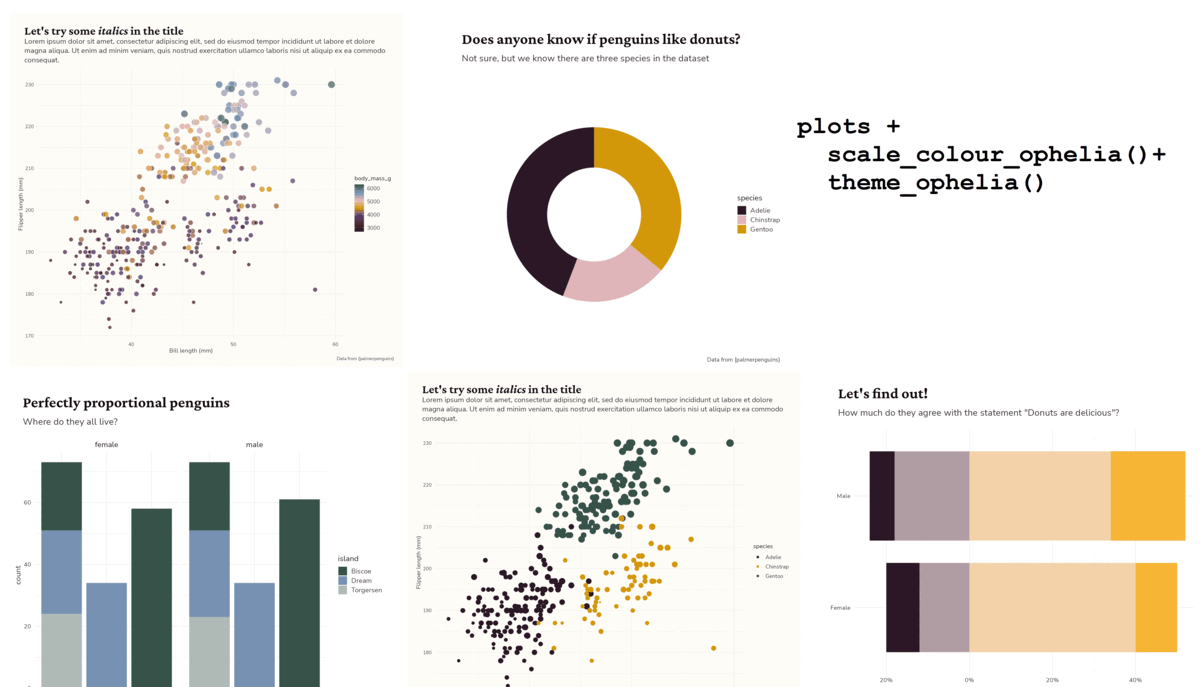
Plot
Helping your future self
Plot + theme_chester_penguins()
Helping your future self
Plot
Helping your future self
Plots + theme_chester_penguins() (or theme_{your-research-group}()?)
Shameless plug alert!
Plots + theme_{your-research-group}()?
Shameless plug alert!
Plots + theme_{your-research-group}()?
<www.cararthompson.com/dataviz-design-systems-explained>
Shameless plug alert!
Plots + theme_{your-research-group}()?
<www.cararthompson.com/dataviz-design-systems-explained>
Annotations
- Main story (range)
- Means within each group
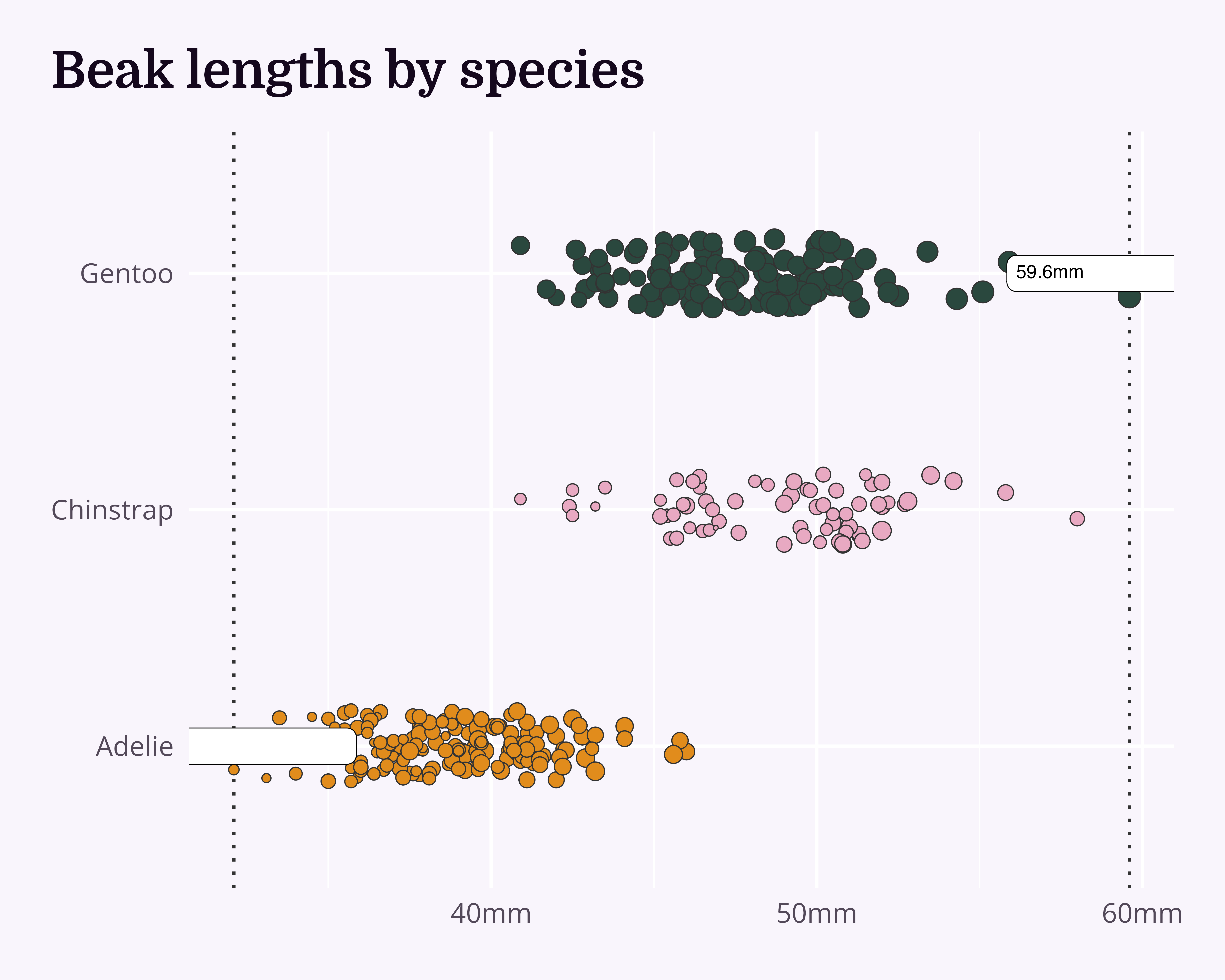
Annotations
Highlight the overall range…
beak_range_df <- penguin_df |>
dplyr::filter(
culmen_length_mm == max(culmen_length_mm, na.rm = TRUE) |
culmen_length_mm == min(culmen_length_mm, na.rm = TRUE)
)
penguin_df |>
ggplot() +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins()Annotations
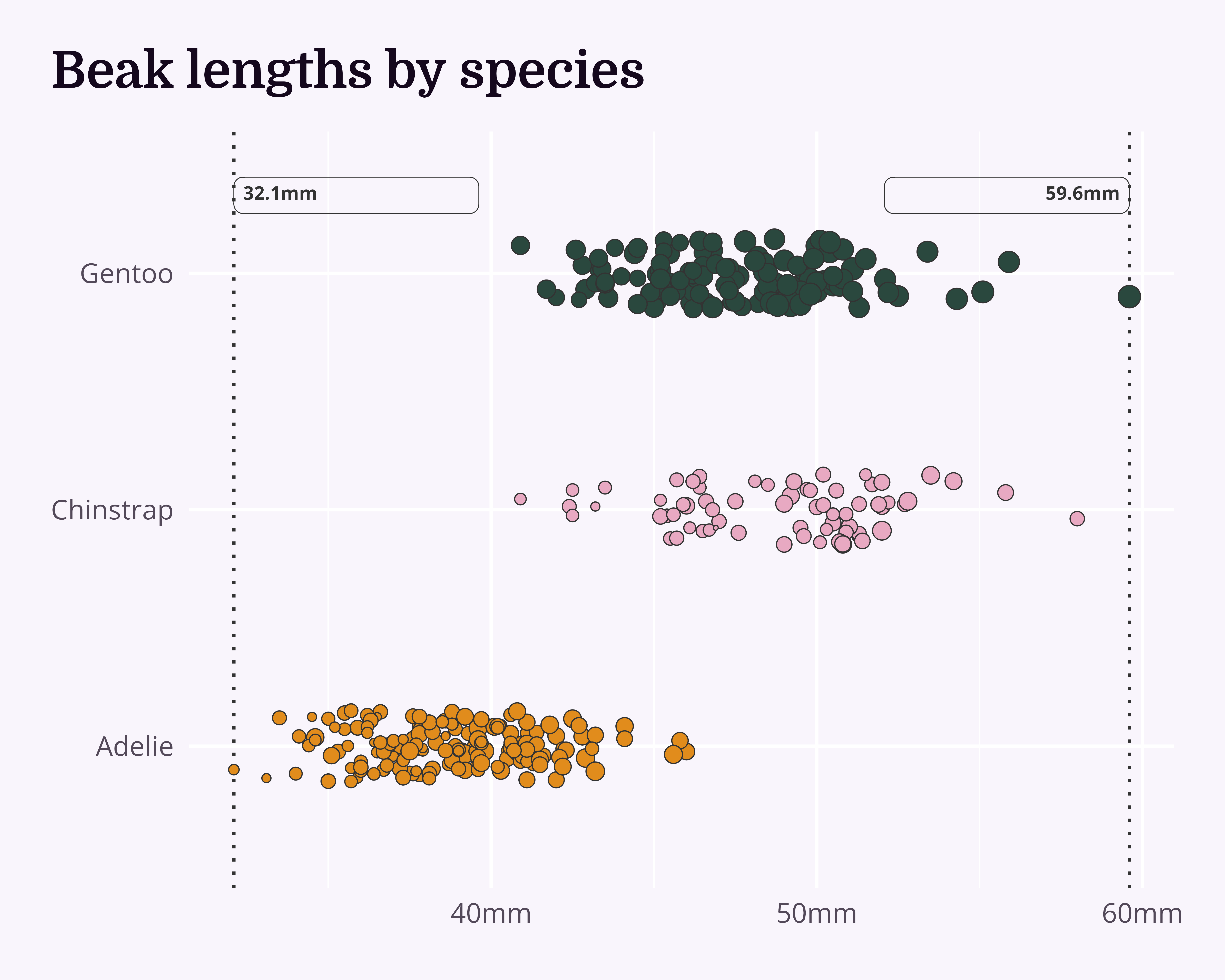
Move to background + change colour
penguin_df |>
ggplot() +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
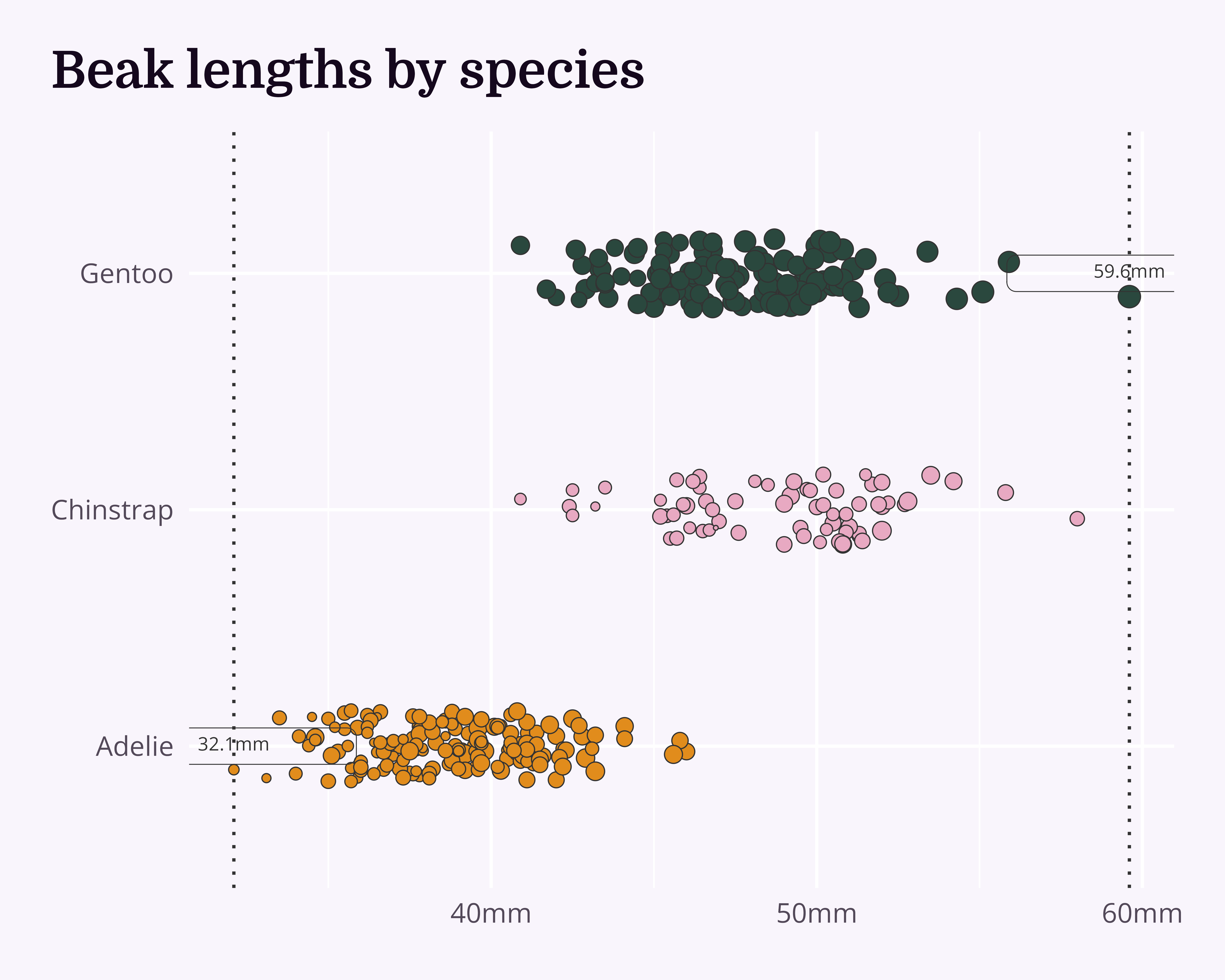
theme_chester_penguins()Annotations
Add labels
penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(label = paste0(culmen_length_mm, "mm"))
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins()Annotations
Add labels
penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(label = paste0(culmen_length_mm, "mm")),
family = "Open Sans",
halign = 0.5,
colour = "#333333",
fill = NA
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
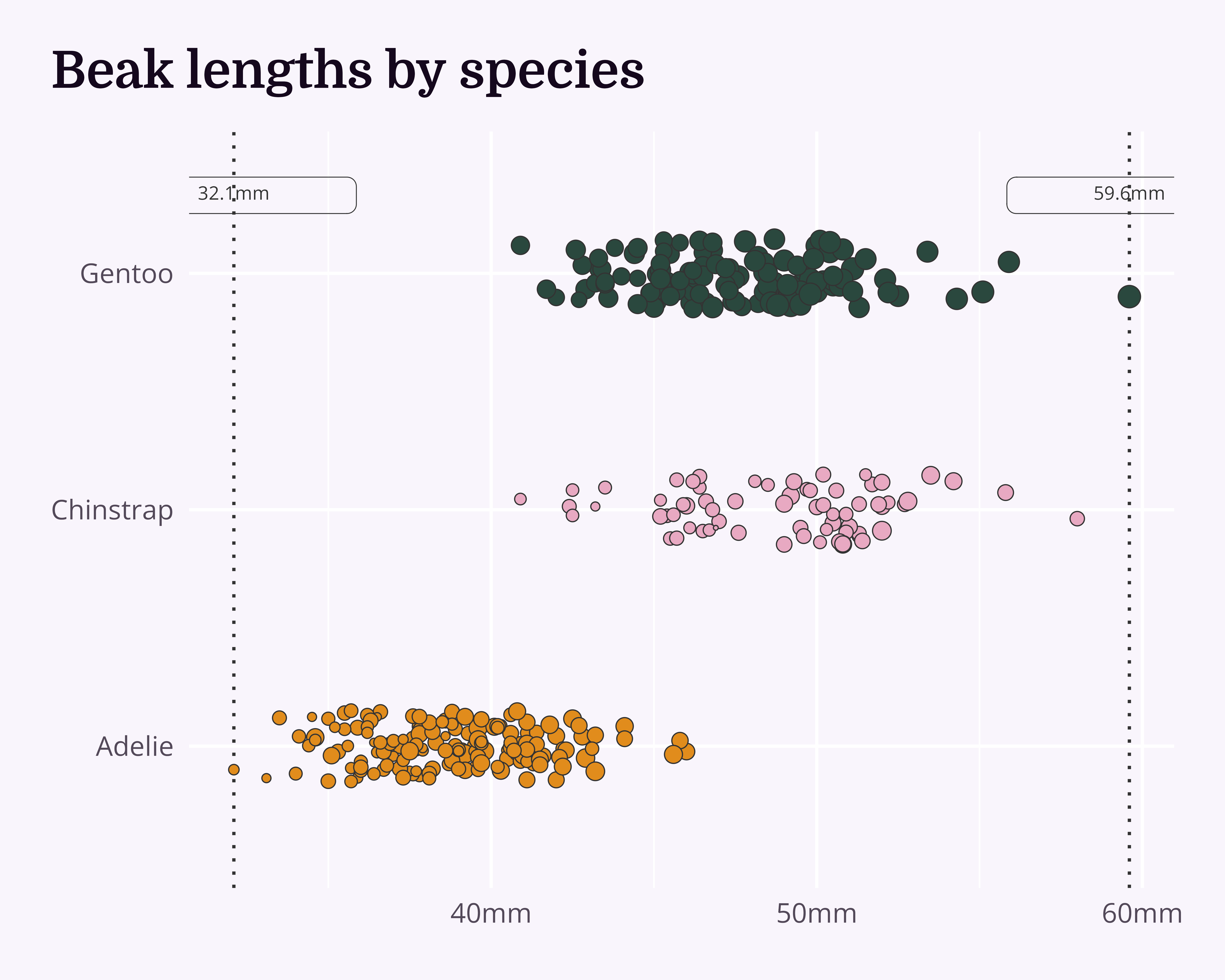
theme_chester_penguins()Annotations
Shift them out of the way of the data
penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(y = max(species), label = paste0(culmen_length_mm, "mm")),
family = "Open Sans",
halign = 0.5,
colour = "#333333",
fill = NA,
nudge_y = 0.33
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins()Annotations
Align them sensibly
penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(
y = max(species),
label = paste0(culmen_length_mm, "mm"),
hjust = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
),
halign = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
)
),
family = "Open Sans",
colour = "#333333",
fill = NA,
nudge_y = 0.33
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins()Annotations
Boldify
penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(
y = max(species),
label = paste0(culmen_length_mm, "mm"),
hjust = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
),
halign = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
)
),
family = "Open Sans",
colour = "#333333",
fontface = "bold",
fill = NA,
nudge_y = 0.33
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins()Annotations
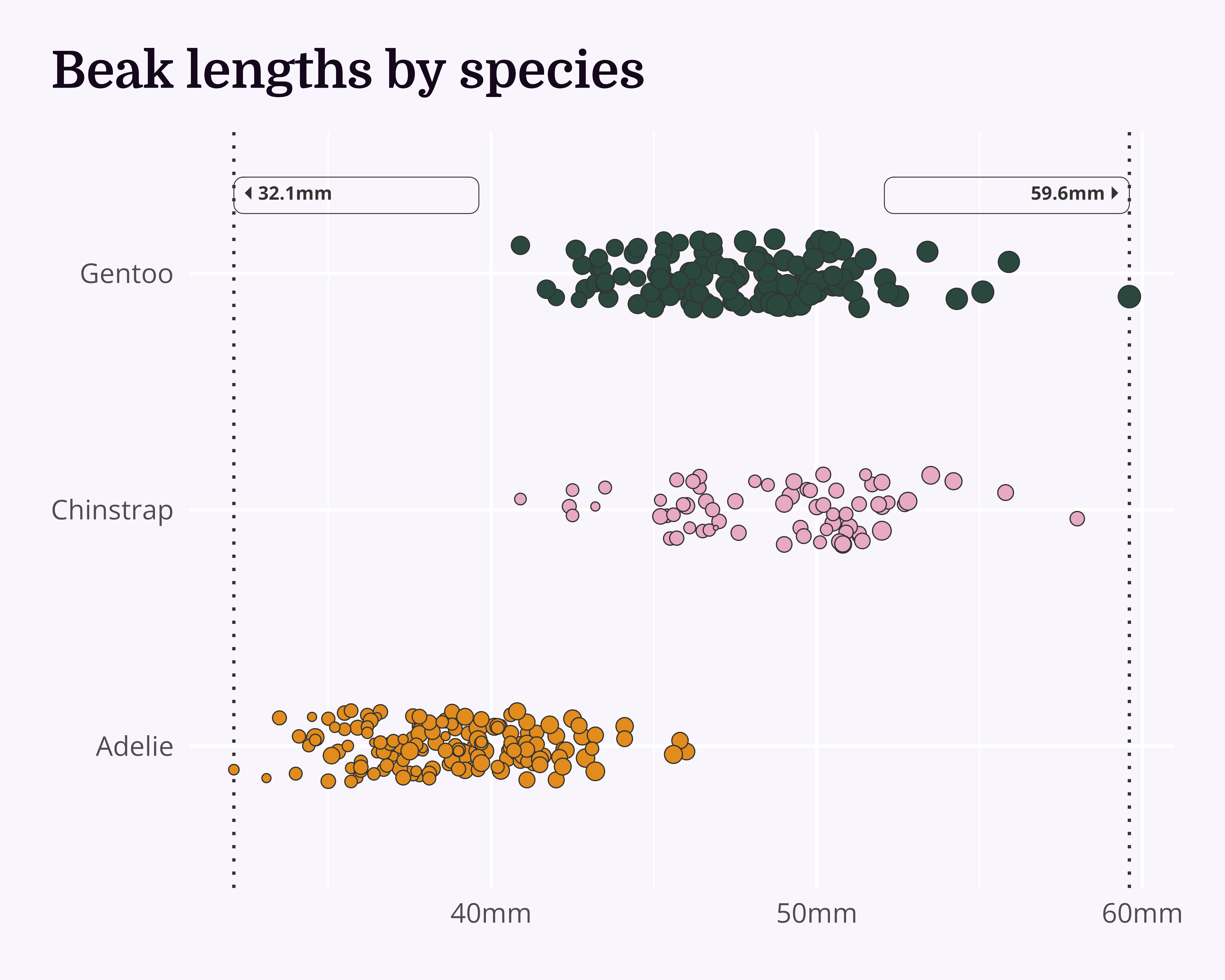
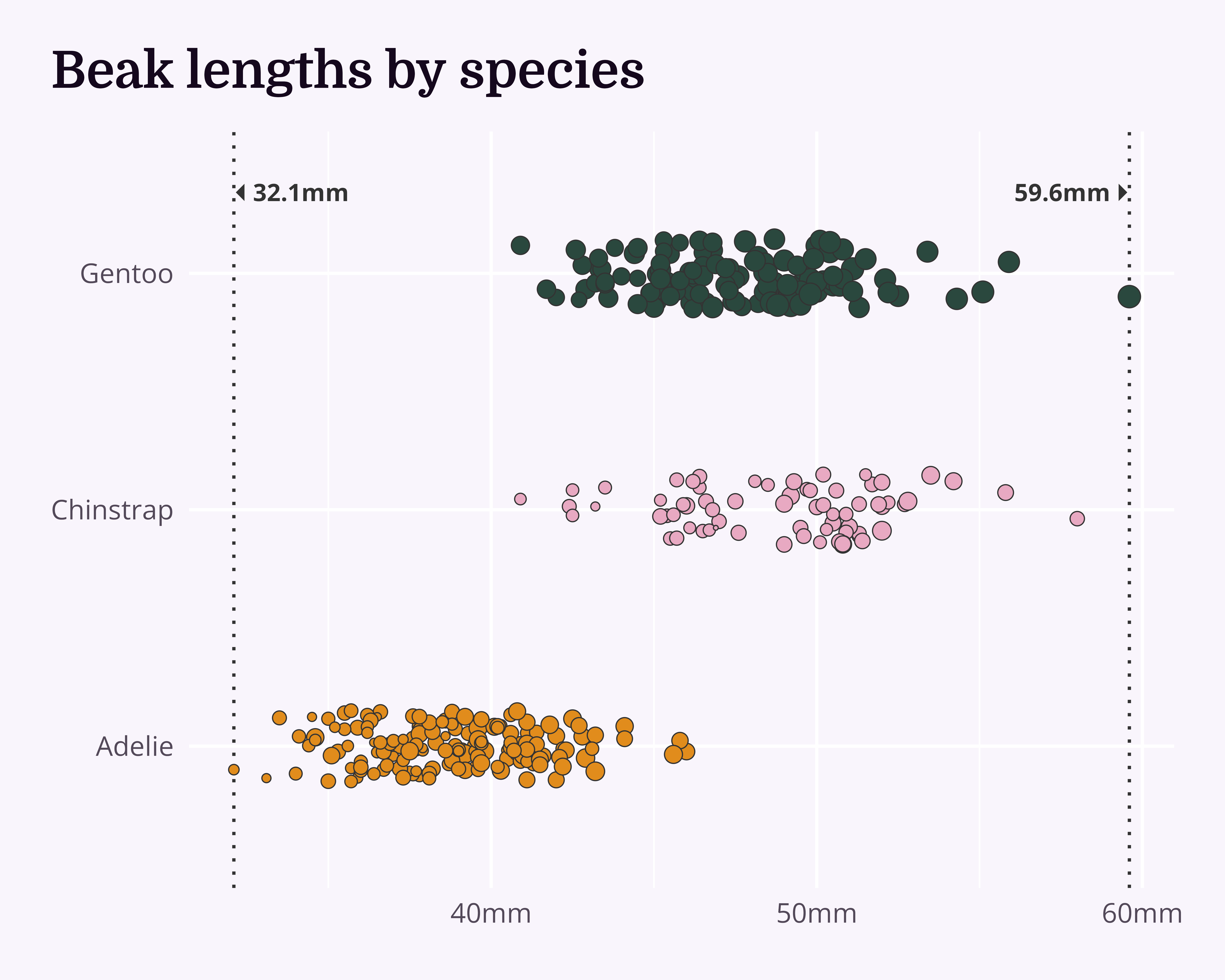
Improve even more!
penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(
y = max(species),
label = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~
paste0("🞀 ", culmen_length_mm, "mm"),
TRUE ~ paste0(culmen_length_mm, "mm", " 🞂")
),
hjust = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
),
halign = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
)
),
family = "Open Sans",
colour = "#333333",
fontface = "bold",
fill = NA,
nudge_y = 0.33
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins()Annotations
Improve even more!
penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(
y = max(species),
label = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~
paste0("🞀 ", culmen_length_mm, "mm"),
TRUE ~ paste0(culmen_length_mm, "mm", " 🞂")
),
hjust = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
),
halign = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
)
),
family = "Open Sans",
colour = "#333333",
fontface = "bold",
fill = NA,
box.padding = unit(0, "pt"),
size = 5,
box.colour = NA,
nudge_y = 0.33
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins()Annotations
And let’s add a bit more data…
beak_means_df <- penguin_df |>
dplyr::group_by(species) |>
dplyr::summarise(mean_length = mean(culmen_length_mm, na.rm = TRUE))
penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_segment(
data = beak_means_df,
aes(x = mean_length, xend = mean_length, y = -Inf, yend = species),
linetype = 3
) +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(
y = max(species),
label = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~
paste0("🞀 ", culmen_length_mm, "mm"),
TRUE ~ paste0(culmen_length_mm, "mm", " 🞂")
),
hjust = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
),
halign = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
)
),
family = "Open Sans",
colour = "#333333",
fontface = "bold",
fill = NA,
box.padding = unit(0, "pt"),
size = 5,
box.colour = NA,
nudge_y = 0.33
) +
ggtext::geom_textbox(
data = beak_means_df,
aes(
x = mean_length,
y = species,
label = paste0(
species,
" mean<br>**",
janitor::round_half_up(mean_length),
"mm**"
)
),
hjust = 0,
nudge_y = -0.4,
box.colour = NA,
family = "Open Sans",
colour = "#333333",
fill = "#f9f5fc"
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins()Annotations
And we can get rid of the y axis!
penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_segment(
data = beak_means_df,
aes(x = mean_length, xend = mean_length, y = -Inf, yend = species),
linetype = 3
) +
geom_jitter(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(
y = max(species),
label = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~
paste0("🞀 ", culmen_length_mm, "mm"),
TRUE ~ paste0(culmen_length_mm, "mm", " 🞂")
),
hjust = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
),
halign = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
)
),
family = "Open Sans",
colour = "#333333",
fontface = "bold",
fill = NA,
box.padding = unit(0, "pt"),
size = 5,
box.colour = NA,
nudge_y = 0.33
) +
ggtext::geom_textbox(
data = beak_means_df,
aes(
x = mean_length,
y = species,
label = paste0(
species,
" mean<br>**",
janitor::round_half_up(mean_length),
"mm**"
)
),
hjust = 0,
nudge_y = -0.4,
box.colour = NA,
family = "Open Sans",
colour = "#333333",
fill = "#f9f5fc"
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins() +
theme(axis.text.y = element_blank())Can we make it interactive?
It’s easier than you think!
Can we make it interactive?
interactive_plot <- penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_segment(
data = beak_means_df,
aes(x = mean_length, xend = mean_length, y = -Inf, yend = species),
linetype = 3
) +
ggiraph::geom_jitter_interactive(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g,
tooltip = paste0("<b>", individual_id, "</b> from ", island)
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(
y = max(species),
label = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~
paste0("🞀 ", culmen_length_mm, "mm"),
TRUE ~ paste0(culmen_length_mm, "mm", " 🞂")
),
hjust = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
),
halign = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
)
),
family = "Open Sans",
colour = "#333333",
fontface = "bold",
fill = NA,
box.padding = unit(0, "pt"),
size = 5,
box.colour = NA,
nudge_y = 0.33
) +
ggtext::geom_textbox(
data = beak_means_df,
aes(
x = mean_length,
y = species,
label = paste0(
species,
" mean<br>**",
janitor::round_half_up(mean_length),
"mm**"
)
),
hjust = 0,
nudge_y = -0.4,
box.colour = NA,
family = "Open Sans",
colour = "#333333",
fill = "#f9f5fc"
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins() +
theme(axis.text.y = element_blank())
ggiraph::girafe(ggobj = interactive_plot)Can we make it interactive?
Let’s style that tooltip!
“Actually, could you just…?”
We want to introduce the penguins in batches
Let’s turn this into a function!
The full code…
penguin_df <- palmerpenguins::penguins_raw |>
janitor::clean_names() |>
dplyr::filter(!is.na(culmen_length_mm))
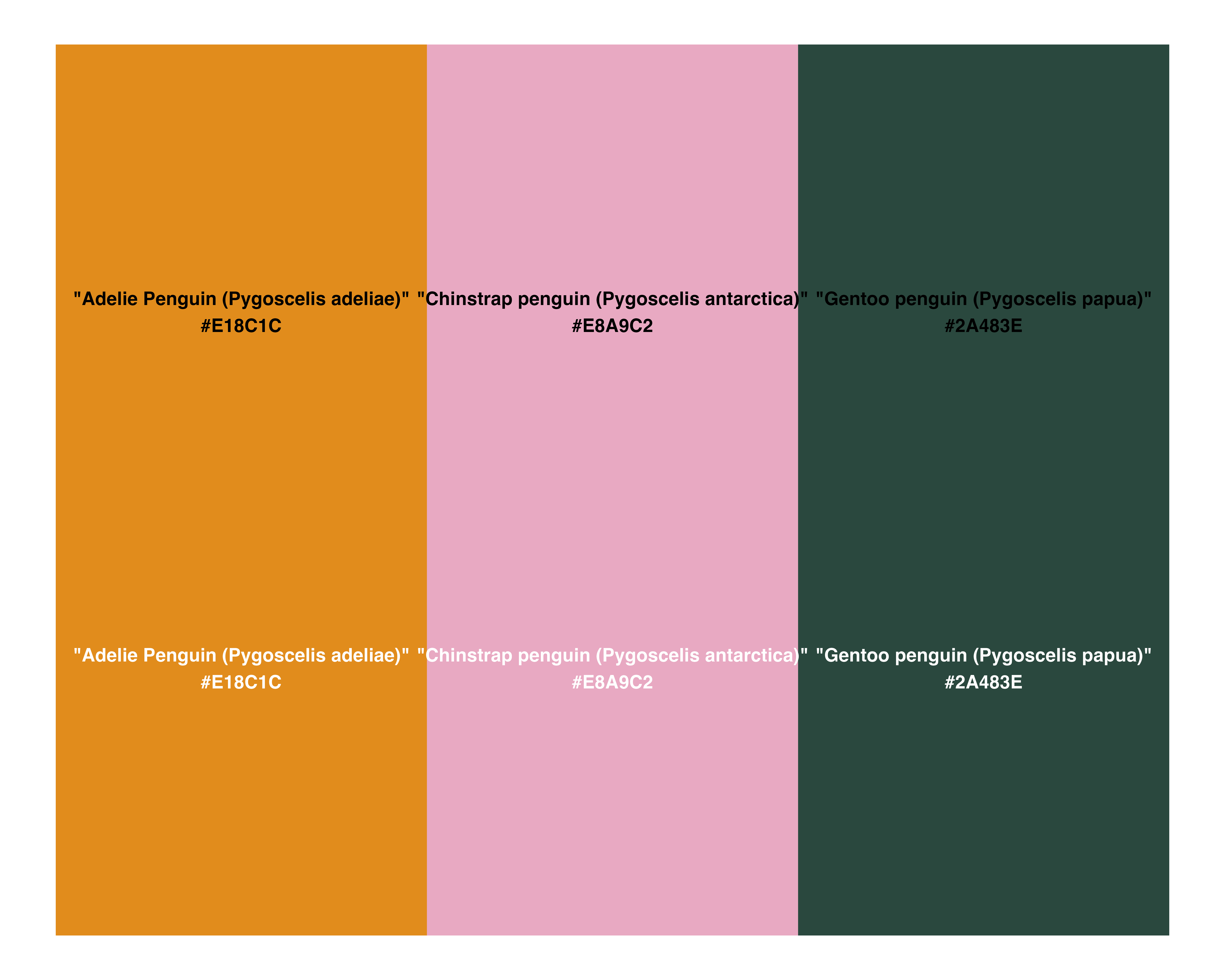
penguin_colours <- c(
"Adelie Penguin (Pygoscelis adeliae)" = "#E18C1C",
"Chinstrap penguin (Pygoscelis antarctica)" = "#E8A9C2",
"Gentoo penguin (Pygoscelis papua)" = "#2A483E"
)
beak_means_df <- penguin_df |>
dplyr::group_by(species) |>
dplyr::summarise(mean_length = mean(culmen_length_mm, na.rm = TRUE))
beak_range_df <- penguin_df |>
dplyr::filter(
culmen_length_mm == max(culmen_length_mm, na.rm = TRUE) |
culmen_length_mm == min(culmen_length_mm, na.rm = TRUE)
)
interactive_plot <- penguin_df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_segment(
data = beak_means_df,
aes(x = mean_length, xend = mean_length, y = -Inf, yend = species),
linetype = 3
) +
ggiraph::geom_jitter_interactive(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g,
tooltip = paste0("<b>", individual_id, "</b> from ", island)
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(
y = max(species),
label = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~
paste0("🞀 ", culmen_length_mm, "mm"),
TRUE ~ paste0(culmen_length_mm, "mm", " 🞂")
),
hjust = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
),
halign = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
)
),
family = "Open Sans",
colour = "#333333",
fontface = "bold",
fill = NA,
box.padding = unit(0, "pt"),
size = 5,
box.colour = NA,
nudge_y = 0.33
) +
ggtext::geom_textbox(
data = beak_means_df,
aes(
x = mean_length,
y = species,
label = paste0(
species,
" mean<br>**",
janitor::round_half_up(mean_length),
"mm**"
)
),
hjust = 0,
nudge_y = -0.4,
box.colour = NA,
family = "Open Sans",
colour = "#333333",
fill = "#f9f5fc"
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = penguin_colours) +
scale_x_continuous(label = function(x) paste0(x, "mm")) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins() +
theme(axis.text.y = element_blank())
ggiraph::girafe(ggobj = interactive_plot)Let’s turn this into a function!
One technicality and two design decisions
penguin_df <- palmerpenguins::penguins_raw |>
janitor::clean_names() |>
dplyr::filter(!is.na(culmen_length_mm))
penguin_colours <- c(
"Adelie Penguin (Pygoscelis adeliae)" = "#E18C1C",
"Chinstrap penguin (Pygoscelis antarctica)" = "#E8A9C2",
"Gentoo penguin (Pygoscelis papua)" = "#2A483E"
)
make_beak_off_plot <- function(
df = penguin_df,
palette = penguin_colours) {
beak_means_df <- df |>
dplyr::group_by(species) |>
dplyr::summarise(mean_length = mean(culmen_length_mm, na.rm = TRUE))
beak_range_df <- df |>
dplyr::filter(
culmen_length_mm == max(culmen_length_mm, na.rm = TRUE) |
culmen_length_mm == min(culmen_length_mm, na.rm = TRUE)
)
interactive_plot <- df |>
ggplot(aes(x = culmen_length_mm, y = species)) +
geom_vline(
data = beak_range_df,
aes(xintercept = culmen_length_mm),
linetype = 3,
colour = "#333333"
) +
geom_segment(
data = beak_means_df,
aes(x = mean_length, xend = mean_length, y = -Inf, yend = species),
linetype = 3
) +
ggiraph::geom_jitter_interactive(
aes(
x = culmen_length_mm,
y = species,
fill = species,
size = body_mass_g,
tooltip = paste0("<b>", individual_id, "</b> from ", island)
),
shape = 21,
width = 0,
height = 0.15,
colour = "#333333",
stroke = 0.5
) +
ggtext::geom_textbox(
data = beak_range_df,
aes(
y = max(df$species),
label = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~
paste0("🞀 ", culmen_length_mm, "mm"),
TRUE ~ paste0(culmen_length_mm, "mm", " 🞂")
),
hjust = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
),
halign = dplyr::case_when(
culmen_length_mm == min(culmen_length_mm) ~ 0,
TRUE ~ 1
)
),
family = "Open Sans",
colour = "#333333",
fontface = "bold",
fill = NA,
box.padding = unit(0, "pt"),
size = 5,
box.colour = NA,
nudge_y = 0.33
) +
ggtext::geom_textbox(
data = beak_means_df,
aes(
x = mean_length,
y = species,
label = paste0(
species,
" mean<br>**",
janitor::round_half_up(mean_length),
"mm**"
)
),
hjust = 0,
nudge_y = -0.4,
box.colour = NA,
family = "Open Sans",
colour = "#333333",
fill = "#f9f5fc"
) +
labs(title = "Beak lengths by species") +
scale_fill_manual(values = palette) +
scale_x_continuous(
label = function(x) paste0(x, "mm"),
limits = range(palmerpenguins::penguins_raw$`Culmen Length (mm)`, na.rm = TRUE)
) +
scale_y_discrete(labels = function(x) gsub("(.)( )(.*)", "\\1", x)) +
theme_chester_penguins() +
theme(axis.text.y = element_blank())
ggiraph::girafe(
ggobj = interactive_plot,
options = list(ggiraph::opts_tooltip(
css = "background-color:#333333;color:#f9f5fc;padding:7.5px;letter-spacing:0.025em;line-height:1.3;border-radius:5px;font-family:Open Sans;"
))
)
}Let’s turn this into a function!
Same graph, different data
Let’s turn this into a function!
Same graph, different data
Where next?
Add some bells and whistles to the function
- Make light outline for dark dots
- Rework the x axis
- Conditional textbox alignments…
Package everything up for easy sharing
- Dependencies ✅
- One source of truth 🪄
- It’s easier than you think!
We’ve already come quite far!
Happy datavizing!

cararthompson.com/talkshello@cararthompson.comcararthompson.com/newsletter
cararthompson.com/talks